library("yulab.utils")14 Gallery of Reproducible Examples
14.1 Visualizing pairwise nucleotide sequence distance with a phylogenetic tree
This example reproduces figure 1 of (Chen et al., 2017). It extracts accession numbers from tip labels of the HPV58 tree and calculates pairwise nucleotide sequence distances. The distance matrix is visualized as dot and line plots. This example demonstrates the ability to add multiple layers to a specific panel. As illustrated in Figure Figure 14.1, the geom_facet() function displays sequence distances as a dot plot and then adds a layer of line plot to the same panel, i.e., sequence distance. In addition, the tree in geom_facet() can be fully annotated with multiple layers (clade labels, bootstrap support values, etc.). The source code is modified from the supplemental file of (Yu et al., 2018).
library(TDbook)
library(tibble)
library(tidyr)
library(Biostrings)Loading required package: BiocGenerics
Attaching package: 'BiocGenerics'The following objects are masked from 'package:stats':
IQR, mad, sd, var, xtabsThe following objects are masked from 'package:base':
anyDuplicated, aperm, append, as.data.frame, basename, cbind,
colnames, dirname, do.call, duplicated, eval, evalq, Filter, Find,
get, grep, grepl, intersect, is.unsorted, lapply, Map, mapply,
match, mget, order, paste, pmax, pmax.int, pmin, pmin.int,
Position, rank, rbind, Reduce, rownames, sapply, saveRDS, setdiff,
table, tapply, union, unique, unsplit, which.max, which.minLoading required package: S4VectorsLoading required package: stats4
Attaching package: 'S4Vectors'The following object is masked from 'package:tidyr':
expandThe following object is masked from 'package:utils':
findMatchesThe following objects are masked from 'package:base':
expand.grid, I, unnameLoading required package: IRangesLoading required package: XVectorLoading required package: GenomeInfoDb
Attaching package: 'Biostrings'The following object is masked from 'package:base':
strsplitlibrary(treeio)treeio v1.35.0 Learn more at https://yulab-smu.top/contribution-tree-data/
Please cite:
LG Wang, TTY Lam, S Xu, Z Dai, L Zhou, T Feng, P Guo, CW Dunn, BR
Jones, T Bradley, H Zhu, Y Guan, Y Jiang, G Yu. treeio: an R package
for phylogenetic tree input and output with richly annotated and
associated data. Molecular Biology and Evolution. 2020, 37(2):599-603.
doi: 10.1093/molbev/msz240
Attaching package: 'treeio'The following object is masked from 'package:Biostrings':
masklibrary(ggplot2)
library(ggtree)ggtree v4.1.2 Learn more at https://yulab-smu.top/contribution-tree-data/
Please cite:
Guangchuang Yu. Data Integration, Manipulation and Visualization of
Phylogenetic Trees (1st edition). Chapman and Hall/CRC. 2022,
doi:10.1201/9781003279242, ISBN: 9781032233574
Attaching package: 'ggtree'The following object is masked from 'package:Biostrings':
collapseThe following object is masked from 'package:IRanges':
collapseThe following object is masked from 'package:S4Vectors':
expandThe following object is masked from 'package:tidyr':
expand# loaded from TDbook package
tree <- tree_HPV58
clade <- c(A3 = 92, A1 = 94, A2 = 108, B1 = 156,
B2 = 159, C = 163, D1 = 173, D2 = 176)
tree <- groupClade(tree, clade)
cols <- c(A1 = "#EC762F", A2 = "#CA6629", A3 = "#894418", B1 = "#0923FA",
B2 = "#020D87", C = "#000000", D1 = "#9ACD32",D2 = "#08630A")
## visualize the tree with tip labels and tree scale
p <- ggtree(tree, aes(color = group), ladderize = FALSE) %>%
rotate(rootnode(tree)) +
geom_tiplab(aes(label = paste0("italic('", label, "')")),
parse = TRUE, size = 2.5) +
geom_treescale(x = 0, y = 1, width = 0.002) +
scale_color_manual(values = c(cols, "black"),
na.value = "black", name = "Lineage",
breaks = c("A1", "A2", "A3", "B1", "B2", "C", "D1", "D2")) +
guides(color = guide_legend(override.aes = list(size = 5, shape = 15))) +
theme_tree2(legend.position = c(.1, .88))Warning: Using `size` aesthetic for lines was deprecated in ggplot2 3.4.0.
ℹ Please use `linewidth` instead.
ℹ The deprecated feature was likely used in the ggtree package.
Please report the issue at <https://github.com/YuLab-SMU/ggtree/issues>.## Optional
## add labels for monophyletic (A, C and D) and paraphyletic (B) groups
dat <- tibble(node = c(94, 108, 131, 92, 156, 159, 163, 173, 176,172),
name = c("A1", "A2", "A3", "A", "B1",
"B2", "C", "D1", "D2", "D"),
offset = c(0.003, 0.003, 0.003, 0.00315, 0.003,
0.003, 0.0031, 0.003, 0.003, 0.00315),
offset.text = c(-.001, -.001, -.001, 0.0002, -.001,
-.001, 0.0002, -.001, -.001, 0.0002),
barsize = c(1.2, 1.2, 1.2, 2, 1.2, 1.2, 3.2, 1.2, 1.2, 2),
extend = list(c(0, 0.5), 0.5, c(0.5, 0), 0, c(0, 0.5),
c(0.5, 0), 0, c(0, 0.5), c(0.5, 0), 0)
) %>%
dplyr::group_split(barsize)
p <- p +
geom_cladelab(
data = dat[[1]],
mapping = aes(
node = node,
label = name,
color = group,
offset = offset,
offset.text = offset.text,
extend = extend
),
barsize = 1.2,
fontface = 3,
align = TRUE
) +
geom_cladelab(
data = dat[[2]],
mapping = aes(
node = node,
label = name,
offset = offset,
offset.text =offset.text,
extend = extend
),
barcolor = "darkgrey",
textcolor = "darkgrey",
barsize = 2,
fontsize = 5,
fontface = 3,
align = TRUE
) +
geom_cladelab(
data = dat[[3]],
mapping = aes(
node = node,
label = name,
offset = offset,
offset.text = offset.text,
extend = extend
),
barcolor = "darkgrey",
textcolor = "darkgrey",
barsize = 3.2,
fontsize = 5,
fontface = 3,
align = TRUE
) +
geom_strip(65, 71, "italic(B)", color = "darkgrey",
offset = 0.00315, align = TRUE, offset.text = 0.0002,
barsize = 2, fontsize = 5, parse = TRUE)
## Optional
## display support values
p <- p + geom_nodelab(aes(subset = (node == 92), label = "*"),
color = "black", nudge_x = -.001, nudge_y = 1) +
geom_nodelab(aes(subset = (node == 155), label = "*"),
color = "black", nudge_x = -.0003, nudge_y = -1) +
geom_nodelab(aes(subset = (node == 158), label = "95/92/1.00"),
color = "black", nudge_x = -0.0001,
nudge_y = -1, hjust = 1) +
geom_nodelab(aes(subset = (node == 162), label = "98/97/1.00"),
color = "black", nudge_x = -0.0001,
nudge_y = -1, hjust = 1) +
geom_nodelab(aes(subset = (node == 172), label = "*"),
color = "black", nudge_x = -.0003, nudge_y = -1) ## extract accession numbers from tip labels
tl <- tree$tip.label
acc <- sub("\\w+\\|", "", tl)
names(tl) <- acc
## read sequences from GenBank directly into R
tipseq <- seqmagick::ncbi_fa_read(acc)
## align the sequences using muscle
tipseq_aln <- muscle::muscle(tipseq)
MUSCLE v3.8.31 by Robert C. Edgar
http://www.drive5.com/muscle
This software is donated to the public domain.
Please cite: Edgar, R.C. Nucleic Acids Res 32(5), 1792-97.
file429f4137593b 90 seqs, max length 7863, avg length 7826
1304 MB(8%)00:00:00 Iter 1 0.02% K-mer dist pass 1
1304 MB(8%)00:00:00 Iter 1 12.23% K-mer dist pass 1
1304 MB(8%)00:00:00 Iter 1 24.44% K-mer dist pass 1
1304 MB(8%)00:00:00 Iter 1 36.65% K-mer dist pass 1
1304 MB(8%)00:00:00 Iter 1 48.86% K-mer dist pass 1
1304 MB(8%)00:00:00 Iter 1 61.07% K-mer dist pass 1
1304 MB(8%)00:00:00 Iter 1 73.28% K-mer dist pass 1
1304 MB(8%)00:00:00 Iter 1 85.49% K-mer dist pass 1
1304 MB(8%)00:00:00 Iter 1 97.70% K-mer dist pass 1
1304 MB(8%)00:00:00 Iter 1 100.00% K-mer dist pass 1
1304 MB(8%)00:00:00 Iter 1 0.02% K-mer dist pass 2
1304 MB(8%)00:00:00 Iter 1 12.23% K-mer dist pass 2
1304 MB(8%)00:00:00 Iter 1 24.44% K-mer dist pass 2
1304 MB(8%)00:00:00 Iter 1 36.65% K-mer dist pass 2
1304 MB(8%)00:00:00 Iter 1 48.86% K-mer dist pass 2
1304 MB(8%)00:00:00 Iter 1 61.07% K-mer dist pass 2
1304 MB(8%)00:00:00 Iter 1 73.28% K-mer dist pass 2
1304 MB(8%)00:00:00 Iter 1 85.49% K-mer dist pass 2
1304 MB(8%)00:00:00 Iter 1 97.70% K-mer dist pass 2
1304 MB(8%)00:00:00 Iter 1 100.00% K-mer dist pass 2
1318 MB(8%)00:00:00 Iter 1 1.12% Align node
1394 MB(9%)00:00:01 Iter 1 2.25% Align node
1405 MB(9%)00:00:02 Iter 1 3.37% Align node
1408 MB(9%)00:00:02 Iter 1 4.49% Align node
1415 MB(9%)00:00:03 Iter 1 5.62% Align node
1422 MB(9%)00:00:03 Iter 1 6.74% Align node
1428 MB(9%)00:00:04 Iter 1 7.87% Align node
1435 MB(9%)00:00:05 Iter 1 8.99% Align node
1447 MB(9%)00:00:05 Iter 1 10.11% Align node
1453 MB(9%)00:00:06 Iter 1 11.24% Align node
1456 MB(9%)00:00:07 Iter 1 12.36% Align node
1458 MB(9%)00:00:07 Iter 1 13.48% Align node
1460 MB(9%)00:00:08 Iter 1 14.61% Align node
1463 MB(9%)00:00:09 Iter 1 15.73% Align node
1465 MB(9%)00:00:09 Iter 1 16.85% Align node
1468 MB(9%)00:00:10 Iter 1 17.98% Align node
1470 MB(9%)00:00:10 Iter 1 19.10% Align node
1472 MB(9%)00:00:11 Iter 1 20.22% Align node
1475 MB(9%)00:00:12 Iter 1 21.35% Align node
1477 MB(9%)00:00:12 Iter 1 22.47% Align node
1479 MB(9%)00:00:13 Iter 1 23.60% Align node
1482 MB(9%)00:00:14 Iter 1 24.72% Align node
1484 MB(9%)00:00:14 Iter 1 25.84% Align node
1495 MB(9%)00:00:15 Iter 1 26.97% Align node
1509 MB(9%)00:00:16 Iter 1 28.09% Align node
1511 MB(9%)00:00:16 Iter 1 29.21% Align node
1520 MB(9%)00:00:17 Iter 1 30.34% Align node
1523 MB(9%)00:00:17 Iter 1 31.46% Align node
1525 MB(9%)00:00:18 Iter 1 32.58% Align node
1528 MB(9%)00:00:19 Iter 1 33.71% Align node
1530 MB(9%)00:00:19 Iter 1 34.83% Align node
1532 MB(9%)00:00:20 Iter 1 35.96% Align node
1535 MB(9%)00:00:21 Iter 1 37.08% Align node
1550 MB(9%)00:00:21 Iter 1 38.20% Align node
1555 MB(9%)00:00:22 Iter 1 39.33% Align node
1562 MB(10%)00:00:23 Iter 1 40.45% Align node
1571 MB(10%)00:00:23 Iter 1 41.57% Align node
1576 MB(10%)00:00:24 Iter 1 42.70% Align node
1578 MB(10%)00:00:25 Iter 1 43.82% Align node
1580 MB(10%)00:00:25 Iter 1 44.94% Align node
1583 MB(10%)00:00:26 Iter 1 46.07% Align node
1585 MB(10%)00:00:26 Iter 1 47.19% Align node
1587 MB(10%)00:00:27 Iter 1 48.31% Align node
1590 MB(10%)00:00:28 Iter 1 49.44% Align node
1592 MB(10%)00:00:28 Iter 1 50.56% Align node
1595 MB(10%)00:00:29 Iter 1 51.69% Align node
1597 MB(10%)00:00:30 Iter 1 52.81% Align node
1604 MB(10%)00:00:30 Iter 1 53.93% Align node
1622 MB(10%)00:00:31 Iter 1 55.06% Align node
1624 MB(10%)00:00:32 Iter 1 56.18% Align node
1627 MB(10%)00:00:32 Iter 1 57.30% Align node
1629 MB(10%)00:00:33 Iter 1 58.43% Align node
1631 MB(10%)00:00:33 Iter 1 59.55% Align node
1634 MB(10%)00:00:34 Iter 1 60.67% Align node
1649 MB(10%)00:00:35 Iter 1 61.80% Align node
1652 MB(10%)00:00:35 Iter 1 62.92% Align node
1654 MB(10%)00:00:36 Iter 1 64.04% Align node
1657 MB(10%)00:00:37 Iter 1 65.17% Align node
1659 MB(10%)00:00:37 Iter 1 66.29% Align node
1661 MB(10%)00:00:38 Iter 1 67.42% Align node
1664 MB(10%)00:00:39 Iter 1 68.54% Align node
1666 MB(10%)00:00:39 Iter 1 69.66% Align node
1668 MB(10%)00:00:40 Iter 1 70.79% Align node
1682 MB(10%)00:00:41 Iter 1 71.91% Align node
1684 MB(10%)00:00:41 Iter 1 73.03% Align node
1687 MB(10%)00:00:42 Iter 1 74.16% Align node
1689 MB(10%)00:00:42 Iter 1 75.28% Align node
1700 MB(10%)00:00:43 Iter 1 76.40% Align node
1703 MB(10%)00:00:44 Iter 1 77.53% Align node
1705 MB(10%)00:00:44 Iter 1 78.65% Align node
1707 MB(10%)00:00:45 Iter 1 79.78% Align node
1716 MB(10%)00:00:46 Iter 1 80.90% Align node
1719 MB(10%)00:00:46 Iter 1 82.02% Align node
1728 MB(11%)00:00:47 Iter 1 83.15% Align node
1739 MB(11%)00:00:48 Iter 1 84.27% Align node
1742 MB(11%)00:00:48 Iter 1 85.39% Align node
1744 MB(11%)00:00:49 Iter 1 86.52% Align node
1746 MB(11%)00:00:49 Iter 1 87.64% Align node
1749 MB(11%)00:00:50 Iter 1 88.76% Align node
1751 MB(11%)00:00:51 Iter 1 89.89% Align node
1760 MB(11%)00:00:51 Iter 1 91.01% Align node
1763 MB(11%)00:00:52 Iter 1 92.13% Align node
1769 MB(11%)00:00:53 Iter 1 93.26% Align node
1776 MB(11%)00:00:53 Iter 1 94.38% Align node
1779 MB(11%)00:00:54 Iter 1 95.51% Align node
1781 MB(11%)00:00:55 Iter 1 96.63% Align node
1783 MB(11%)00:00:55 Iter 1 97.75% Align node
1786 MB(11%)00:00:56 Iter 1 98.88% Align node
1788 MB(11%)00:00:57 Iter 1 100.00% Align node
1790 MB(11%)00:00:57 Iter 1 100.00% Align node
1790 MB(11%)00:00:57 Iter 1 1.11% Root alignment
1791 MB(11%)00:00:57 Iter 1 2.22% Root alignment
1791 MB(11%)00:00:57 Iter 1 3.33% Root alignment
1791 MB(11%)00:00:57 Iter 1 4.44% Root alignment
1791 MB(11%)00:00:57 Iter 1 5.56% Root alignment
1791 MB(11%)00:00:57 Iter 1 6.67% Root alignment
1791 MB(11%)00:00:57 Iter 1 7.78% Root alignment
1791 MB(11%)00:00:57 Iter 1 8.89% Root alignment
1791 MB(11%)00:00:57 Iter 1 10.00% Root alignment
1791 MB(11%)00:00:57 Iter 1 11.11% Root alignment
1791 MB(11%)00:00:57 Iter 1 12.22% Root alignment
1791 MB(11%)00:00:57 Iter 1 13.33% Root alignment
1791 MB(11%)00:00:57 Iter 1 14.44% Root alignment
1791 MB(11%)00:00:57 Iter 1 15.56% Root alignment
1791 MB(11%)00:00:57 Iter 1 16.67% Root alignment
1791 MB(11%)00:00:57 Iter 1 17.78% Root alignment
1791 MB(11%)00:00:57 Iter 1 18.89% Root alignment
1791 MB(11%)00:00:57 Iter 1 20.00% Root alignment
1791 MB(11%)00:00:57 Iter 1 21.11% Root alignment
1791 MB(11%)00:00:57 Iter 1 22.22% Root alignment
1791 MB(11%)00:00:57 Iter 1 23.33% Root alignment
1791 MB(11%)00:00:57 Iter 1 24.44% Root alignment
1791 MB(11%)00:00:57 Iter 1 25.56% Root alignment
1791 MB(11%)00:00:57 Iter 1 26.67% Root alignment
1791 MB(11%)00:00:57 Iter 1 27.78% Root alignment
1791 MB(11%)00:00:57 Iter 1 28.89% Root alignment
1791 MB(11%)00:00:57 Iter 1 30.00% Root alignment
1791 MB(11%)00:00:57 Iter 1 31.11% Root alignment
1791 MB(11%)00:00:57 Iter 1 32.22% Root alignment
1791 MB(11%)00:00:57 Iter 1 33.33% Root alignment
1791 MB(11%)00:00:57 Iter 1 34.44% Root alignment
1791 MB(11%)00:00:57 Iter 1 35.56% Root alignment
1791 MB(11%)00:00:57 Iter 1 36.67% Root alignment
1791 MB(11%)00:00:57 Iter 1 37.78% Root alignment
1791 MB(11%)00:00:57 Iter 1 38.89% Root alignment
1791 MB(11%)00:00:57 Iter 1 40.00% Root alignment
1791 MB(11%)00:00:57 Iter 1 41.11% Root alignment
1791 MB(11%)00:00:57 Iter 1 42.22% Root alignment
1791 MB(11%)00:00:57 Iter 1 43.33% Root alignment
1791 MB(11%)00:00:57 Iter 1 44.44% Root alignment
1791 MB(11%)00:00:57 Iter 1 45.56% Root alignment
1791 MB(11%)00:00:57 Iter 1 46.67% Root alignment
1791 MB(11%)00:00:57 Iter 1 47.78% Root alignment
1791 MB(11%)00:00:57 Iter 1 48.89% Root alignment
1791 MB(11%)00:00:57 Iter 1 50.00% Root alignment
1791 MB(11%)00:00:57 Iter 1 51.11% Root alignment
1791 MB(11%)00:00:57 Iter 1 52.22% Root alignment
1791 MB(11%)00:00:57 Iter 1 53.33% Root alignment
1791 MB(11%)00:00:57 Iter 1 54.44% Root alignment
1791 MB(11%)00:00:57 Iter 1 55.56% Root alignment
1791 MB(11%)00:00:57 Iter 1 56.67% Root alignment
1791 MB(11%)00:00:57 Iter 1 57.78% Root alignment
1791 MB(11%)00:00:57 Iter 1 58.89% Root alignment
1791 MB(11%)00:00:57 Iter 1 60.00% Root alignment
1791 MB(11%)00:00:57 Iter 1 61.11% Root alignment
1791 MB(11%)00:00:57 Iter 1 62.22% Root alignment
1791 MB(11%)00:00:57 Iter 1 63.33% Root alignment
1791 MB(11%)00:00:57 Iter 1 64.44% Root alignment
1791 MB(11%)00:00:57 Iter 1 65.56% Root alignment
1791 MB(11%)00:00:57 Iter 1 66.67% Root alignment
1791 MB(11%)00:00:57 Iter 1 67.78% Root alignment
1791 MB(11%)00:00:57 Iter 1 68.89% Root alignment
1791 MB(11%)00:00:57 Iter 1 70.00% Root alignment
1791 MB(11%)00:00:57 Iter 1 71.11% Root alignment
1791 MB(11%)00:00:57 Iter 1 72.22% Root alignment
1791 MB(11%)00:00:57 Iter 1 73.33% Root alignment
1791 MB(11%)00:00:57 Iter 1 74.44% Root alignment
1791 MB(11%)00:00:57 Iter 1 75.56% Root alignment
1791 MB(11%)00:00:57 Iter 1 76.67% Root alignment
1791 MB(11%)00:00:57 Iter 1 77.78% Root alignment
1791 MB(11%)00:00:57 Iter 1 78.89% Root alignment
1791 MB(11%)00:00:57 Iter 1 80.00% Root alignment
1791 MB(11%)00:00:57 Iter 1 81.11% Root alignment
1791 MB(11%)00:00:57 Iter 1 82.22% Root alignment
1791 MB(11%)00:00:57 Iter 1 83.33% Root alignment
1791 MB(11%)00:00:57 Iter 1 84.44% Root alignment
1791 MB(11%)00:00:57 Iter 1 85.56% Root alignment
1791 MB(11%)00:00:57 Iter 1 86.67% Root alignment
1791 MB(11%)00:00:57 Iter 1 87.78% Root alignment
1791 MB(11%)00:00:57 Iter 1 88.89% Root alignment
1791 MB(11%)00:00:57 Iter 1 90.00% Root alignment
1791 MB(11%)00:00:57 Iter 1 91.11% Root alignment
1791 MB(11%)00:00:57 Iter 1 92.22% Root alignment
1791 MB(11%)00:00:57 Iter 1 93.33% Root alignment
1791 MB(11%)00:00:57 Iter 1 94.44% Root alignment
1791 MB(11%)00:00:57 Iter 1 95.56% Root alignment
1791 MB(11%)00:00:57 Iter 1 96.67% Root alignment
1791 MB(11%)00:00:57 Iter 1 97.78% Root alignment
1791 MB(11%)00:00:57 Iter 1 98.89% Root alignment
1791 MB(11%)00:00:57 Iter 1 100.00% Root alignment
1791 MB(11%)00:00:57 Iter 1 100.00% Root alignment
1791 MB(11%)00:00:57 Iter 2 1.14% Refine tree
1793 MB(11%)00:00:58 Iter 2 2.27% Refine tree
1793 MB(11%)00:00:59 Iter 2 3.41% Refine tree
1793 MB(11%)00:00:59 Iter 2 4.55% Refine tree
1793 MB(11%)00:01:00 Iter 2 5.68% Refine tree
1793 MB(11%)00:01:01 Iter 2 6.82% Refine tree
1793 MB(11%)00:01:01 Iter 2 7.95% Refine tree
1793 MB(11%)00:01:02 Iter 2 9.09% Refine tree
1793 MB(11%)00:01:02 Iter 2 10.23% Refine tree
1793 MB(11%)00:01:03 Iter 2 11.36% Refine tree
1793 MB(11%)00:01:04 Iter 2 12.50% Refine tree
1793 MB(11%)00:01:04 Iter 2 13.64% Refine tree
1793 MB(11%)00:01:05 Iter 2 14.77% Refine tree
1793 MB(11%)00:01:06 Iter 2 15.91% Refine tree
1793 MB(11%)00:01:06 Iter 2 17.05% Refine tree
1793 MB(11%)00:01:07 Iter 2 18.18% Refine tree
1793 MB(11%)00:01:08 Iter 2 19.32% Refine tree
1793 MB(11%)00:01:08 Iter 2 20.45% Refine tree
1793 MB(11%)00:01:09 Iter 2 21.59% Refine tree
1793 MB(11%)00:01:09 Iter 2 22.73% Refine tree
1793 MB(11%)00:01:10 Iter 2 23.86% Refine tree
1793 MB(11%)00:01:11 Iter 2 25.00% Refine tree
1793 MB(11%)00:01:11 Iter 2 26.14% Refine tree
1793 MB(11%)00:01:12 Iter 2 27.27% Refine tree
1793 MB(11%)00:01:13 Iter 2 28.41% Refine tree
1793 MB(11%)00:01:13 Iter 2 29.55% Refine tree
1793 MB(11%)00:01:14 Iter 2 30.68% Refine tree
1793 MB(11%)00:01:15 Iter 2 31.82% Refine tree
1793 MB(11%)00:01:15 Iter 2 32.95% Refine tree
1793 MB(11%)00:01:16 Iter 2 34.09% Refine tree
1793 MB(11%)00:01:16 Iter 2 35.23% Refine tree
1793 MB(11%)00:01:17 Iter 2 36.36% Refine tree
1793 MB(11%)00:01:18 Iter 2 37.50% Refine tree
1793 MB(11%)00:01:18 Iter 2 38.64% Refine tree
1793 MB(11%)00:01:19 Iter 2 39.77% Refine tree
1793 MB(11%)00:01:20 Iter 2 40.91% Refine tree
1793 MB(11%)00:01:20 Iter 2 42.05% Refine tree
1793 MB(11%)00:01:21 Iter 2 43.18% Refine tree
1793 MB(11%)00:01:22 Iter 2 44.32% Refine tree
1793 MB(11%)00:01:22 Iter 2 45.45% Refine tree
1793 MB(11%)00:01:23 Iter 2 46.59% Refine tree
1793 MB(11%)00:01:24 Iter 2 47.73% Refine tree
1793 MB(11%)00:01:24 Iter 2 48.86% Refine tree
1793 MB(11%)00:01:25 Iter 2 50.00% Refine tree
1793 MB(11%)00:01:25 Iter 2 51.14% Refine tree
1793 MB(11%)00:01:26 Iter 2 52.27% Refine tree
1793 MB(11%)00:01:27 Iter 2 53.41% Refine tree
1793 MB(11%)00:01:27 Iter 2 54.55% Refine tree
1793 MB(11%)00:01:28 Iter 2 100.00% Refine tree
1793 MB(11%)00:01:28 Iter 2 1.11% Root alignment
1793 MB(11%)00:01:28 Iter 2 2.22% Root alignment
1793 MB(11%)00:01:28 Iter 2 3.33% Root alignment
1793 MB(11%)00:01:28 Iter 2 4.44% Root alignment
1793 MB(11%)00:01:28 Iter 2 5.56% Root alignment
1793 MB(11%)00:01:28 Iter 2 6.67% Root alignment
1793 MB(11%)00:01:28 Iter 2 7.78% Root alignment
1793 MB(11%)00:01:28 Iter 2 8.89% Root alignment
1793 MB(11%)00:01:28 Iter 2 10.00% Root alignment
1793 MB(11%)00:01:28 Iter 2 11.11% Root alignment
1793 MB(11%)00:01:28 Iter 2 12.22% Root alignment
1793 MB(11%)00:01:28 Iter 2 13.33% Root alignment
1793 MB(11%)00:01:28 Iter 2 14.44% Root alignment
1793 MB(11%)00:01:28 Iter 2 15.56% Root alignment
1793 MB(11%)00:01:28 Iter 2 16.67% Root alignment
1793 MB(11%)00:01:28 Iter 2 17.78% Root alignment
1793 MB(11%)00:01:28 Iter 2 18.89% Root alignment
1793 MB(11%)00:01:28 Iter 2 20.00% Root alignment
1793 MB(11%)00:01:28 Iter 2 21.11% Root alignment
1793 MB(11%)00:01:28 Iter 2 22.22% Root alignment
1793 MB(11%)00:01:28 Iter 2 23.33% Root alignment
1793 MB(11%)00:01:28 Iter 2 24.44% Root alignment
1793 MB(11%)00:01:28 Iter 2 25.56% Root alignment
1793 MB(11%)00:01:28 Iter 2 26.67% Root alignment
1793 MB(11%)00:01:28 Iter 2 27.78% Root alignment
1793 MB(11%)00:01:28 Iter 2 28.89% Root alignment
1793 MB(11%)00:01:28 Iter 2 30.00% Root alignment
1793 MB(11%)00:01:28 Iter 2 31.11% Root alignment
1793 MB(11%)00:01:28 Iter 2 32.22% Root alignment
1793 MB(11%)00:01:28 Iter 2 33.33% Root alignment
1793 MB(11%)00:01:28 Iter 2 34.44% Root alignment
1793 MB(11%)00:01:28 Iter 2 35.56% Root alignment
1793 MB(11%)00:01:28 Iter 2 36.67% Root alignment
1793 MB(11%)00:01:28 Iter 2 37.78% Root alignment
1793 MB(11%)00:01:28 Iter 2 38.89% Root alignment
1793 MB(11%)00:01:28 Iter 2 40.00% Root alignment
1793 MB(11%)00:01:28 Iter 2 41.11% Root alignment
1793 MB(11%)00:01:28 Iter 2 42.22% Root alignment
1793 MB(11%)00:01:28 Iter 2 43.33% Root alignment
1793 MB(11%)00:01:28 Iter 2 44.44% Root alignment
1793 MB(11%)00:01:28 Iter 2 45.56% Root alignment
1793 MB(11%)00:01:28 Iter 2 46.67% Root alignment
1793 MB(11%)00:01:28 Iter 2 47.78% Root alignment
1793 MB(11%)00:01:28 Iter 2 48.89% Root alignment
1793 MB(11%)00:01:28 Iter 2 50.00% Root alignment
1793 MB(11%)00:01:28 Iter 2 51.11% Root alignment
1793 MB(11%)00:01:28 Iter 2 52.22% Root alignment
1793 MB(11%)00:01:28 Iter 2 53.33% Root alignment
1793 MB(11%)00:01:28 Iter 2 54.44% Root alignment
1793 MB(11%)00:01:28 Iter 2 55.56% Root alignment
1793 MB(11%)00:01:28 Iter 2 56.67% Root alignment
1793 MB(11%)00:01:28 Iter 2 57.78% Root alignment
1793 MB(11%)00:01:28 Iter 2 58.89% Root alignment
1793 MB(11%)00:01:28 Iter 2 60.00% Root alignment
1793 MB(11%)00:01:28 Iter 2 61.11% Root alignment
1793 MB(11%)00:01:28 Iter 2 62.22% Root alignment
1793 MB(11%)00:01:28 Iter 2 63.33% Root alignment
1793 MB(11%)00:01:28 Iter 2 64.44% Root alignment
1793 MB(11%)00:01:28 Iter 2 65.56% Root alignment
1793 MB(11%)00:01:28 Iter 2 66.67% Root alignment
1793 MB(11%)00:01:28 Iter 2 67.78% Root alignment
1793 MB(11%)00:01:28 Iter 2 68.89% Root alignment
1793 MB(11%)00:01:28 Iter 2 70.00% Root alignment
1793 MB(11%)00:01:28 Iter 2 71.11% Root alignment
1793 MB(11%)00:01:28 Iter 2 72.22% Root alignment
1793 MB(11%)00:01:28 Iter 2 73.33% Root alignment
1793 MB(11%)00:01:28 Iter 2 74.44% Root alignment
1793 MB(11%)00:01:28 Iter 2 75.56% Root alignment
1793 MB(11%)00:01:28 Iter 2 76.67% Root alignment
1793 MB(11%)00:01:28 Iter 2 77.78% Root alignment
1793 MB(11%)00:01:28 Iter 2 78.89% Root alignment
1793 MB(11%)00:01:28 Iter 2 80.00% Root alignment
1793 MB(11%)00:01:28 Iter 2 81.11% Root alignment
1793 MB(11%)00:01:28 Iter 2 82.22% Root alignment
1793 MB(11%)00:01:28 Iter 2 83.33% Root alignment
1793 MB(11%)00:01:28 Iter 2 84.44% Root alignment
1793 MB(11%)00:01:28 Iter 2 85.56% Root alignment
1793 MB(11%)00:01:28 Iter 2 86.67% Root alignment
1793 MB(11%)00:01:28 Iter 2 87.78% Root alignment
1793 MB(11%)00:01:28 Iter 2 88.89% Root alignment
1793 MB(11%)00:01:28 Iter 2 90.00% Root alignment
1793 MB(11%)00:01:28 Iter 2 91.11% Root alignment
1793 MB(11%)00:01:28 Iter 2 92.22% Root alignment
1793 MB(11%)00:01:28 Iter 2 93.33% Root alignment
1793 MB(11%)00:01:28 Iter 2 94.44% Root alignment
1793 MB(11%)00:01:28 Iter 2 95.56% Root alignment
1793 MB(11%)00:01:28 Iter 2 96.67% Root alignment
1793 MB(11%)00:01:28 Iter 2 97.78% Root alignment
1793 MB(11%)00:01:28 Iter 2 98.89% Root alignment
1793 MB(11%)00:01:28 Iter 2 100.00% Root alignment
1793 MB(11%)00:01:28 Iter 2 100.00% Root alignment
1793 MB(11%)00:01:28 Iter 2 100.00% Root alignment
1793 MB(11%)00:01:29 Iter 3 1.13% Refine biparts
1793 MB(11%)00:01:30 Iter 3 1.69% Refine biparts
1793 MB(11%)00:01:30 Iter 3 2.26% Refine biparts
1793 MB(11%)00:01:31 Iter 3 2.82% Refine biparts
1793 MB(11%)00:01:32 Iter 3 3.39% Refine biparts
1793 MB(11%)00:01:32 Iter 3 3.95% Refine biparts
1793 MB(11%)00:01:33 Iter 3 4.52% Refine biparts
1793 MB(11%)00:01:34 Iter 3 5.08% Refine biparts
1793 MB(11%)00:01:34 Iter 3 5.65% Refine biparts
1793 MB(11%)00:01:35 Iter 3 6.21% Refine biparts
1793 MB(11%)00:01:36 Iter 3 6.78% Refine biparts
1793 MB(11%)00:01:36 Iter 3 7.34% Refine biparts
1793 MB(11%)00:01:37 Iter 3 7.91% Refine biparts
1793 MB(11%)00:01:38 Iter 3 8.47% Refine biparts
1793 MB(11%)00:01:38 Iter 3 9.04% Refine biparts
1793 MB(11%)00:01:39 Iter 3 9.60% Refine biparts
1793 MB(11%)00:01:40 Iter 3 10.17% Refine biparts
1793 MB(11%)00:01:40 Iter 3 10.73% Refine biparts
1793 MB(11%)00:01:41 Iter 3 11.30% Refine biparts
1793 MB(11%)00:01:42 Iter 3 11.86% Refine biparts
1793 MB(11%)00:01:42 Iter 3 12.43% Refine biparts
1793 MB(11%)00:01:43 Iter 3 12.99% Refine biparts
1793 MB(11%)00:01:44 Iter 3 13.56% Refine biparts
1793 MB(11%)00:01:44 Iter 3 14.12% Refine biparts
1793 MB(11%)00:01:45 Iter 3 14.69% Refine biparts
1793 MB(11%)00:01:46 Iter 3 15.25% Refine biparts
1793 MB(11%)00:01:47 Iter 3 15.82% Refine biparts
1793 MB(11%)00:01:47 Iter 3 16.38% Refine biparts
1793 MB(11%)00:01:48 Iter 3 16.95% Refine biparts
1793 MB(11%)00:01:49 Iter 3 17.51% Refine biparts
1793 MB(11%)00:01:49 Iter 3 18.08% Refine biparts
1793 MB(11%)00:01:50 Iter 3 18.64% Refine biparts
1793 MB(11%)00:01:51 Iter 3 19.21% Refine biparts
1793 MB(11%)00:01:51 Iter 3 19.77% Refine biparts
1793 MB(11%)00:01:52 Iter 3 20.34% Refine biparts
1793 MB(11%)00:01:53 Iter 3 20.90% Refine biparts
1793 MB(11%)00:01:53 Iter 3 21.47% Refine biparts
1793 MB(11%)00:01:54 Iter 3 22.03% Refine biparts
1793 MB(11%)00:01:55 Iter 3 22.60% Refine biparts
1793 MB(11%)00:01:56 Iter 3 23.16% Refine biparts
1793 MB(11%)00:01:56 Iter 3 23.73% Refine biparts
1793 MB(11%)00:01:57 Iter 3 24.29% Refine biparts
1793 MB(11%)00:01:58 Iter 3 24.86% Refine biparts
1793 MB(11%)00:01:58 Iter 3 25.42% Refine biparts
1793 MB(11%)00:01:59 Iter 3 25.99% Refine biparts
1793 MB(11%)00:02:00 Iter 3 26.55% Refine biparts
1793 MB(11%)00:02:00 Iter 3 27.12% Refine biparts
1793 MB(11%)00:02:01 Iter 3 27.68% Refine biparts
1793 MB(11%)00:02:02 Iter 3 28.25% Refine biparts
1793 MB(11%)00:02:02 Iter 3 28.81% Refine biparts
1793 MB(11%)00:02:03 Iter 3 29.38% Refine biparts
1793 MB(11%)00:02:04 Iter 3 29.94% Refine biparts
1793 MB(11%)00:02:05 Iter 3 30.51% Refine biparts
1793 MB(11%)00:02:05 Iter 3 31.07% Refine biparts
1793 MB(11%)00:02:06 Iter 3 31.64% Refine biparts
1793 MB(11%)00:02:07 Iter 3 32.20% Refine biparts
1793 MB(11%)00:02:07 Iter 3 32.77% Refine biparts
1793 MB(11%)00:02:08 Iter 3 33.33% Refine biparts
1793 MB(11%)00:02:09 Iter 3 33.90% Refine biparts
1793 MB(11%)00:02:09 Iter 3 34.46% Refine biparts
1793 MB(11%)00:02:10 Iter 3 35.03% Refine biparts
1793 MB(11%)00:02:11 Iter 3 35.59% Refine biparts
1793 MB(11%)00:02:11 Iter 3 36.16% Refine biparts
1793 MB(11%)00:02:12 Iter 3 36.72% Refine biparts
1793 MB(11%)00:02:13 Iter 3 37.29% Refine biparts
1793 MB(11%)00:02:14 Iter 3 37.85% Refine biparts
1793 MB(11%)00:02:14 Iter 3 38.42% Refine biparts
1793 MB(11%)00:02:15 Iter 3 38.98% Refine biparts
1793 MB(11%)00:02:16 Iter 3 39.55% Refine biparts
1793 MB(11%)00:02:16 Iter 3 40.11% Refine biparts
1793 MB(11%)00:02:17 Iter 3 40.68% Refine biparts
1793 MB(11%)00:02:18 Iter 3 41.24% Refine biparts
1793 MB(11%)00:02:19 Iter 3 41.81% Refine biparts
1793 MB(11%)00:02:19 Iter 3 42.37% Refine biparts
1793 MB(11%)00:02:20 Iter 3 42.94% Refine biparts
1793 MB(11%)00:02:21 Iter 3 43.50% Refine biparts
1793 MB(11%)00:02:21 Iter 3 44.07% Refine biparts
1793 MB(11%)00:02:22 Iter 3 44.63% Refine biparts
1793 MB(11%)00:02:23 Iter 3 45.20% Refine biparts
1793 MB(11%)00:02:23 Iter 3 45.76% Refine biparts
1793 MB(11%)00:02:24 Iter 3 46.33% Refine biparts
1793 MB(11%)00:02:25 Iter 3 46.89% Refine biparts
1793 MB(11%)00:02:26 Iter 3 47.46% Refine biparts
1793 MB(11%)00:02:26 Iter 3 48.02% Refine biparts
1793 MB(11%)00:02:27 Iter 3 48.59% Refine biparts
1793 MB(11%)00:02:28 Iter 3 49.15% Refine biparts
1793 MB(11%)00:02:28 Iter 3 49.72% Refine biparts
1793 MB(11%)00:02:29 Iter 3 50.28% Refine biparts
1793 MB(11%)00:02:30 Iter 3 50.85% Refine biparts
1793 MB(11%)00:02:31 Iter 3 51.41% Refine biparts
1793 MB(11%)00:02:31 Iter 3 51.98% Refine biparts
1793 MB(11%)00:02:32 Iter 3 52.54% Refine biparts
1793 MB(11%)00:02:33 Iter 3 53.11% Refine biparts
1793 MB(11%)00:02:33 Iter 3 53.67% Refine biparts
1793 MB(11%)00:02:34 Iter 3 54.24% Refine biparts
1793 MB(11%)00:02:35 Iter 3 54.80% Refine biparts
1793 MB(11%)00:02:35 Iter 3 55.37% Refine biparts
1793 MB(11%)00:02:36 Iter 3 55.93% Refine biparts
1793 MB(11%)00:02:37 Iter 3 56.50% Refine biparts
1793 MB(11%)00:02:37 Iter 3 57.06% Refine biparts
1793 MB(11%)00:02:38 Iter 3 57.63% Refine biparts
1793 MB(11%)00:02:39 Iter 3 58.19% Refine biparts
1793 MB(11%)00:02:39 Iter 3 58.76% Refine biparts
1793 MB(11%)00:02:40 Iter 3 59.32% Refine biparts
1793 MB(11%)00:02:41 Iter 3 59.89% Refine biparts
1793 MB(11%)00:02:41 Iter 3 60.45% Refine biparts
1793 MB(11%)00:02:42 Iter 3 61.02% Refine biparts
1793 MB(11%)00:02:43 Iter 3 61.58% Refine biparts
1793 MB(11%)00:02:43 Iter 3 62.15% Refine biparts
1793 MB(11%)00:02:44 Iter 3 62.71% Refine biparts
1793 MB(11%)00:02:45 Iter 3 63.28% Refine biparts
1793 MB(11%)00:02:46 Iter 3 63.84% Refine biparts
1793 MB(11%)00:02:46 Iter 3 64.41% Refine biparts
1793 MB(11%)00:02:47 Iter 3 64.97% Refine biparts
1793 MB(11%)00:02:48 Iter 3 65.54% Refine biparts
1793 MB(11%)00:02:48 Iter 3 66.10% Refine biparts
1793 MB(11%)00:02:49 Iter 3 66.67% Refine biparts
1793 MB(11%)00:02:50 Iter 3 67.23% Refine biparts
1793 MB(11%)00:02:50 Iter 3 67.80% Refine biparts
1793 MB(11%)00:02:51 Iter 3 68.36% Refine biparts
1793 MB(11%)00:02:52 Iter 3 68.93% Refine biparts
1793 MB(11%)00:02:52 Iter 3 69.49% Refine biparts
1793 MB(11%)00:02:53 Iter 3 70.06% Refine biparts
1793 MB(11%)00:02:54 Iter 3 70.62% Refine biparts
1793 MB(11%)00:02:54 Iter 3 71.19% Refine biparts
1793 MB(11%)00:02:55 Iter 3 71.75% Refine biparts
1793 MB(11%)00:02:56 Iter 3 72.32% Refine biparts
1793 MB(11%)00:02:56 Iter 3 72.88% Refine biparts
1793 MB(11%)00:02:57 Iter 3 73.45% Refine biparts
1793 MB(11%)00:02:58 Iter 3 74.01% Refine biparts
1793 MB(11%)00:02:58 Iter 3 74.58% Refine biparts
1793 MB(11%)00:02:59 Iter 3 75.14% Refine biparts
1793 MB(11%)00:03:00 Iter 3 75.71% Refine biparts
1793 MB(11%)00:03:01 Iter 3 76.27% Refine biparts
1793 MB(11%)00:03:01 Iter 3 76.84% Refine biparts
1793 MB(11%)00:03:02 Iter 3 77.40% Refine biparts
1793 MB(11%)00:03:03 Iter 3 77.97% Refine biparts
1793 MB(11%)00:03:03 Iter 3 78.53% Refine biparts
1793 MB(11%)00:03:04 Iter 3 79.10% Refine biparts
1793 MB(11%)00:03:05 Iter 3 79.66% Refine biparts
1793 MB(11%)00:03:05 Iter 3 80.23% Refine biparts
1793 MB(11%)00:03:06 Iter 3 80.79% Refine biparts
1793 MB(11%)00:03:07 Iter 3 81.36% Refine biparts
1793 MB(11%)00:03:07 Iter 3 81.92% Refine biparts
1793 MB(11%)00:03:08 Iter 3 82.49% Refine biparts
1793 MB(11%)00:03:09 Iter 3 83.05% Refine biparts
1793 MB(11%)00:03:09 Iter 3 83.62% Refine biparts
1793 MB(11%)00:03:10 Iter 3 84.18% Refine biparts
1793 MB(11%)00:03:11 Iter 3 84.75% Refine biparts
1793 MB(11%)00:03:11 Iter 3 85.31% Refine biparts
1793 MB(11%)00:03:12 Iter 3 85.88% Refine biparts
1793 MB(11%)00:03:13 Iter 3 86.44% Refine biparts
1793 MB(11%)00:03:14 Iter 3 87.01% Refine biparts
1793 MB(11%)00:03:14 Iter 3 87.57% Refine biparts
1793 MB(11%)00:03:15 Iter 3 88.14% Refine biparts
1793 MB(11%)00:03:16 Iter 3 88.70% Refine biparts
1793 MB(11%)00:03:16 Iter 3 89.27% Refine biparts
1793 MB(11%)00:03:17 Iter 3 89.83% Refine biparts
1793 MB(11%)00:03:18 Iter 3 90.40% Refine biparts
1793 MB(11%)00:03:18 Iter 3 90.96% Refine biparts
1793 MB(11%)00:03:19 Iter 3 91.53% Refine biparts
1793 MB(11%)00:03:20 Iter 3 92.09% Refine biparts
1793 MB(11%)00:03:20 Iter 3 92.66% Refine biparts
1793 MB(11%)00:03:21 Iter 3 93.22% Refine biparts
1793 MB(11%)00:03:22 Iter 3 93.79% Refine biparts
1793 MB(11%)00:03:23 Iter 3 94.35% Refine biparts
1793 MB(11%)00:03:23 Iter 3 94.92% Refine biparts
1793 MB(11%)00:03:24 Iter 3 95.48% Refine biparts
1793 MB(11%)00:03:25 Iter 3 96.05% Refine biparts
1793 MB(11%)00:03:25 Iter 3 96.61% Refine biparts
1793 MB(11%)00:03:26 Iter 3 97.18% Refine biparts
1793 MB(11%)00:03:27 Iter 3 97.74% Refine biparts
1793 MB(11%)00:03:27 Iter 3 98.31% Refine biparts
1793 MB(11%)00:03:28 Iter 3 98.87% Refine biparts
1793 MB(11%)00:03:29 Iter 3 99.44% Refine biparts
1793 MB(11%)00:03:30 Iter 3 100.00% Refine biparts
1793 MB(11%)00:03:30 Iter 3 100.56% Refine biparts
1793 MB(11%)00:03:30 Iter 3 100.00% Refine biparts
1793 MB(11%)00:03:31 Iter 4 1.13% Refine biparts
1793 MB(11%)00:03:32 Iter 4 1.69% Refine biparts
1793 MB(11%)00:03:32 Iter 4 2.26% Refine biparts
1793 MB(11%)00:03:33 Iter 4 2.82% Refine biparts
1793 MB(11%)00:03:34 Iter 4 3.39% Refine biparts
1793 MB(11%)00:03:35 Iter 4 3.95% Refine biparts
1793 MB(11%)00:03:35 Iter 4 4.52% Refine biparts
1793 MB(11%)00:03:36 Iter 4 5.08% Refine biparts
1793 MB(11%)00:03:37 Iter 4 5.65% Refine biparts
1793 MB(11%)00:03:37 Iter 4 6.21% Refine biparts
1793 MB(11%)00:03:38 Iter 4 6.78% Refine biparts
1793 MB(11%)00:03:39 Iter 4 7.34% Refine biparts
1793 MB(11%)00:03:39 Iter 4 7.91% Refine biparts
1793 MB(11%)00:03:40 Iter 4 8.47% Refine biparts
1793 MB(11%)00:03:41 Iter 4 9.04% Refine biparts
1793 MB(11%)00:03:42 Iter 4 9.60% Refine biparts
1793 MB(11%)00:03:42 Iter 4 10.17% Refine biparts
1793 MB(11%)00:03:43 Iter 4 10.73% Refine biparts
1793 MB(11%)00:03:44 Iter 4 11.30% Refine biparts
1793 MB(11%)00:03:44 Iter 4 11.86% Refine biparts
1793 MB(11%)00:03:45 Iter 4 12.43% Refine biparts
1793 MB(11%)00:03:46 Iter 4 12.99% Refine biparts
1793 MB(11%)00:03:46 Iter 4 13.56% Refine biparts
1793 MB(11%)00:03:47 Iter 4 14.12% Refine biparts
1793 MB(11%)00:03:48 Iter 4 14.69% Refine biparts
1793 MB(11%)00:03:49 Iter 4 15.25% Refine biparts
1793 MB(11%)00:03:49 Iter 4 15.82% Refine biparts
1793 MB(11%)00:03:50 Iter 4 16.38% Refine biparts
1793 MB(11%)00:03:51 Iter 4 16.95% Refine biparts
1793 MB(11%)00:03:51 Iter 4 17.51% Refine biparts
1793 MB(11%)00:03:52 Iter 4 18.08% Refine biparts
1793 MB(11%)00:03:53 Iter 4 18.64% Refine biparts
1793 MB(11%)00:03:53 Iter 4 19.21% Refine biparts
1793 MB(11%)00:03:54 Iter 4 19.77% Refine biparts
1793 MB(11%)00:03:55 Iter 4 20.34% Refine biparts
1793 MB(11%)00:03:55 Iter 4 20.90% Refine biparts
1793 MB(11%)00:03:56 Iter 4 21.47% Refine biparts
1793 MB(11%)00:03:57 Iter 4 22.03% Refine biparts
1793 MB(11%)00:03:58 Iter 4 22.60% Refine biparts
1793 MB(11%)00:03:58 Iter 4 23.16% Refine biparts
1793 MB(11%)00:03:59 Iter 4 23.73% Refine biparts
1793 MB(11%)00:04:00 Iter 4 24.29% Refine biparts
1793 MB(11%)00:04:00 Iter 4 24.86% Refine biparts
1793 MB(11%)00:04:01 Iter 4 25.42% Refine biparts
1793 MB(11%)00:04:02 Iter 4 25.99% Refine biparts
1793 MB(11%)00:04:02 Iter 4 26.55% Refine biparts
1793 MB(11%)00:04:03 Iter 4 27.12% Refine biparts
1793 MB(11%)00:04:04 Iter 4 27.68% Refine biparts
1793 MB(11%)00:04:05 Iter 4 28.25% Refine biparts
1793 MB(11%)00:04:05 Iter 4 28.81% Refine biparts
1793 MB(11%)00:04:06 Iter 4 29.38% Refine biparts
1793 MB(11%)00:04:07 Iter 4 29.94% Refine biparts
1793 MB(11%)00:04:07 Iter 4 30.51% Refine biparts
1793 MB(11%)00:04:08 Iter 4 31.07% Refine biparts
1793 MB(11%)00:04:09 Iter 4 31.64% Refine biparts
1793 MB(11%)00:04:09 Iter 4 32.20% Refine biparts
1793 MB(11%)00:04:10 Iter 4 32.77% Refine biparts
1793 MB(11%)00:04:11 Iter 4 33.33% Refine biparts
1793 MB(11%)00:04:11 Iter 4 33.90% Refine biparts
1793 MB(11%)00:04:12 Iter 4 34.46% Refine biparts
1793 MB(11%)00:04:13 Iter 4 35.03% Refine biparts
1793 MB(11%)00:04:13 Iter 4 35.59% Refine biparts
1793 MB(11%)00:04:14 Iter 4 36.16% Refine biparts
1793 MB(11%)00:04:15 Iter 4 36.72% Refine biparts
1793 MB(11%)00:04:16 Iter 4 37.29% Refine biparts
1793 MB(11%)00:04:16 Iter 4 37.85% Refine biparts
1793 MB(11%)00:04:17 Iter 4 38.42% Refine biparts
1793 MB(11%)00:04:18 Iter 4 38.98% Refine biparts
1793 MB(11%)00:04:18 Iter 4 39.55% Refine biparts
1793 MB(11%)00:04:19 Iter 4 40.11% Refine biparts
1793 MB(11%)00:04:20 Iter 4 40.68% Refine biparts
1793 MB(11%)00:04:20 Iter 4 41.24% Refine biparts
1793 MB(11%)00:04:21 Iter 4 41.81% Refine biparts
1793 MB(11%)00:04:22 Iter 4 42.37% Refine biparts
1793 MB(11%)00:04:22 Iter 4 42.94% Refine biparts
1793 MB(11%)00:04:23 Iter 4 43.50% Refine biparts
1793 MB(11%)00:04:24 Iter 4 44.07% Refine biparts
1793 MB(11%)00:04:24 Iter 4 44.63% Refine biparts
1793 MB(11%)00:04:25 Iter 4 45.20% Refine biparts
1793 MB(11%)00:04:26 Iter 4 45.76% Refine biparts
1793 MB(11%)00:04:26 Iter 4 46.33% Refine biparts
1793 MB(11%)00:04:27 Iter 4 46.89% Refine biparts
1793 MB(11%)00:04:28 Iter 4 47.46% Refine biparts
1793 MB(11%)00:04:28 Iter 4 48.02% Refine biparts
1793 MB(11%)00:04:29 Iter 4 48.59% Refine biparts
1793 MB(11%)00:04:30 Iter 4 49.15% Refine biparts
1793 MB(11%)00:04:31 Iter 4 49.72% Refine biparts
1793 MB(11%)00:04:31 Iter 4 50.28% Refine biparts
1793 MB(11%)00:04:32 Iter 4 50.85% Refine biparts
1793 MB(11%)00:04:33 Iter 4 51.41% Refine biparts
1793 MB(11%)00:04:33 Iter 4 51.98% Refine biparts
1793 MB(11%)00:04:34 Iter 4 52.54% Refine biparts
1793 MB(11%)00:04:35 Iter 4 53.11% Refine biparts
1793 MB(11%)00:04:35 Iter 4 53.67% Refine biparts
1793 MB(11%)00:04:36 Iter 4 54.24% Refine biparts
1793 MB(11%)00:04:37 Iter 4 54.80% Refine biparts
1793 MB(11%)00:04:38 Iter 4 55.37% Refine biparts
1793 MB(11%)00:04:38 Iter 4 55.93% Refine biparts
1793 MB(11%)00:04:39 Iter 4 56.50% Refine biparts
1793 MB(11%)00:04:40 Iter 4 57.06% Refine biparts
1793 MB(11%)00:04:40 Iter 4 57.63% Refine biparts
1793 MB(11%)00:04:41 Iter 4 58.19% Refine biparts
1793 MB(11%)00:04:42 Iter 4 58.76% Refine biparts
1793 MB(11%)00:04:42 Iter 4 59.32% Refine biparts
1793 MB(11%)00:04:43 Iter 4 59.89% Refine biparts
1793 MB(11%)00:04:44 Iter 4 60.45% Refine biparts
1793 MB(11%)00:04:44 Iter 4 61.02% Refine biparts
1793 MB(11%)00:04:45 Iter 4 61.58% Refine biparts
1793 MB(11%)00:04:46 Iter 4 62.15% Refine biparts
1793 MB(11%)00:04:47 Iter 4 62.71% Refine biparts
1793 MB(11%)00:04:47 Iter 4 63.28% Refine biparts
1793 MB(11%)00:04:48 Iter 4 63.84% Refine biparts
1793 MB(11%)00:04:49 Iter 4 64.41% Refine biparts
1793 MB(11%)00:04:49 Iter 4 64.97% Refine biparts
1793 MB(11%)00:04:50 Iter 4 65.54% Refine biparts
1793 MB(11%)00:04:51 Iter 4 66.10% Refine biparts
1793 MB(11%)00:04:51 Iter 4 66.67% Refine biparts
1793 MB(11%)00:04:52 Iter 4 67.23% Refine biparts
1793 MB(11%)00:04:53 Iter 4 67.80% Refine biparts
1793 MB(11%)00:04:53 Iter 4 68.36% Refine biparts
1793 MB(11%)00:04:54 Iter 4 68.93% Refine biparts
1793 MB(11%)00:04:55 Iter 4 69.49% Refine biparts
1793 MB(11%)00:04:55 Iter 4 70.06% Refine biparts
1793 MB(11%)00:04:56 Iter 4 70.62% Refine biparts
1793 MB(11%)00:04:57 Iter 4 71.19% Refine biparts
1793 MB(11%)00:04:57 Iter 4 71.75% Refine biparts
1793 MB(11%)00:04:58 Iter 4 72.32% Refine biparts
1793 MB(11%)00:04:59 Iter 4 72.88% Refine biparts
1793 MB(11%)00:05:00 Iter 4 73.45% Refine biparts
1793 MB(11%)00:05:00 Iter 4 74.01% Refine biparts
1793 MB(11%)00:05:01 Iter 4 74.58% Refine biparts
1793 MB(11%)00:05:02 Iter 4 75.14% Refine biparts
1793 MB(11%)00:05:02 Iter 4 75.71% Refine biparts
1793 MB(11%)00:05:03 Iter 4 76.27% Refine biparts
1793 MB(11%)00:05:04 Iter 4 76.84% Refine biparts
1793 MB(11%)00:05:04 Iter 4 77.40% Refine biparts
1793 MB(11%)00:05:05 Iter 4 77.97% Refine biparts
1793 MB(11%)00:05:06 Iter 4 78.53% Refine biparts
1793 MB(11%)00:05:06 Iter 4 79.10% Refine biparts
1793 MB(11%)00:05:07 Iter 4 79.66% Refine biparts
1793 MB(11%)00:05:08 Iter 4 80.23% Refine biparts
1793 MB(11%)00:05:08 Iter 4 80.79% Refine biparts
1793 MB(11%)00:05:09 Iter 4 81.36% Refine biparts
1793 MB(11%)00:05:10 Iter 4 81.92% Refine biparts
1793 MB(11%)00:05:11 Iter 4 82.49% Refine biparts
1793 MB(11%)00:05:11 Iter 4 83.05% Refine biparts
1793 MB(11%)00:05:12 Iter 4 83.62% Refine biparts
1793 MB(11%)00:05:13 Iter 4 84.18% Refine biparts
1793 MB(11%)00:05:13 Iter 4 84.75% Refine biparts
1793 MB(11%)00:05:14 Iter 4 85.31% Refine biparts
1793 MB(11%)00:05:15 Iter 4 85.88% Refine biparts
1793 MB(11%)00:05:15 Iter 4 86.44% Refine biparts
1793 MB(11%)00:05:16 Iter 4 87.01% Refine biparts
1793 MB(11%)00:05:17 Iter 4 87.57% Refine biparts
1793 MB(11%)00:05:17 Iter 4 88.14% Refine biparts
1793 MB(11%)00:05:18 Iter 4 88.70% Refine biparts
1793 MB(11%)00:05:19 Iter 4 89.27% Refine biparts
1793 MB(11%)00:05:19 Iter 4 89.83% Refine biparts
1793 MB(11%)00:05:20 Iter 4 90.40% Refine biparts
1793 MB(11%)00:05:21 Iter 4 90.96% Refine biparts
1793 MB(11%)00:05:21 Iter 4 91.53% Refine biparts
1793 MB(11%)00:05:22 Iter 4 92.09% Refine biparts
1793 MB(11%)00:05:23 Iter 4 92.66% Refine biparts
1793 MB(11%)00:05:24 Iter 4 93.22% Refine biparts
1793 MB(11%)00:05:24 Iter 4 93.79% Refine biparts
1793 MB(11%)00:05:25 Iter 4 94.35% Refine biparts
1793 MB(11%)00:05:26 Iter 4 94.92% Refine biparts
1793 MB(11%)00:05:26 Iter 4 95.48% Refine biparts
1793 MB(11%)00:05:27 Iter 4 96.05% Refine biparts
1793 MB(11%)00:05:28 Iter 4 96.61% Refine biparts
1793 MB(11%)00:05:28 Iter 4 97.18% Refine biparts
1793 MB(11%)00:05:29 Iter 4 97.74% Refine biparts
1793 MB(11%)00:05:30 Iter 4 98.31% Refine biparts
1793 MB(11%)00:05:30 Iter 4 98.87% Refine biparts
1793 MB(11%)00:05:31 Iter 4 99.44% Refine biparts
1793 MB(11%)00:05:32 Iter 4 100.00% Refine biparts
1793 MB(11%)00:05:32 Iter 4 100.56% Refine biparts
1793 MB(11%)00:05:32 Iter 4 100.00% Refine bipartstipseq_aln <- DNAStringSet(tipseq_aln)## extract accession numbers from tip labels
tl <- tree$tip.label
acc <- sub("\\w+\\|", "", tl)
names(tl) <- acc
## writeXStringSet(tipseq_aln, file = "data/HPV58_aln.fas")
#tipseq_aln <- readDNAStringSet("data/HPV58_aln.fas")
tipseq_aln <- TDbook::dna_HPV58_aln %>%
as.character %>%
lapply(., paste0, collapse = "") %>%
unlist() %>%
Biostrings::DNAStringSet()## calculate pairwise hamming distances among sequences
tipseq_dist <- pwalign::stringDist(tipseq_aln, method = "hamming")
## calculate the percentage of differences
tipseq_d <- as.matrix(tipseq_dist) / width(tipseq_aln[1]) * 100
## convert the matrix to a tidy data frame for facet_plot
dd <- as_tibble(tipseq_d)
dd$seq1 <- rownames(tipseq_d)
td <- gather(dd,seq2, dist, -seq1)
td$seq1 <- tl[td$seq1]
td$seq2 <- tl[td$seq2]
g <- p$data$group
names(g) <- p$data$label
td$clade <- g[td$seq2]
## visualize the sequence differences using dot plot and line plot
## and align the sequence difference plot to the tree using facet_plot
p2 <- p + geom_facet(panel = "Sequence Distance",
data = td, geom = geom_point, alpha = .6,
mapping = aes(x = dist, color = clade, shape = clade)) +
geom_facet(panel = "Sequence Distance",
data = td, geom = geom_path, alpha = .6,
mapping=aes(x = dist, group = seq2, color = clade)) +
scale_shape_manual(values = 1:8, guide = FALSE)
print(p2)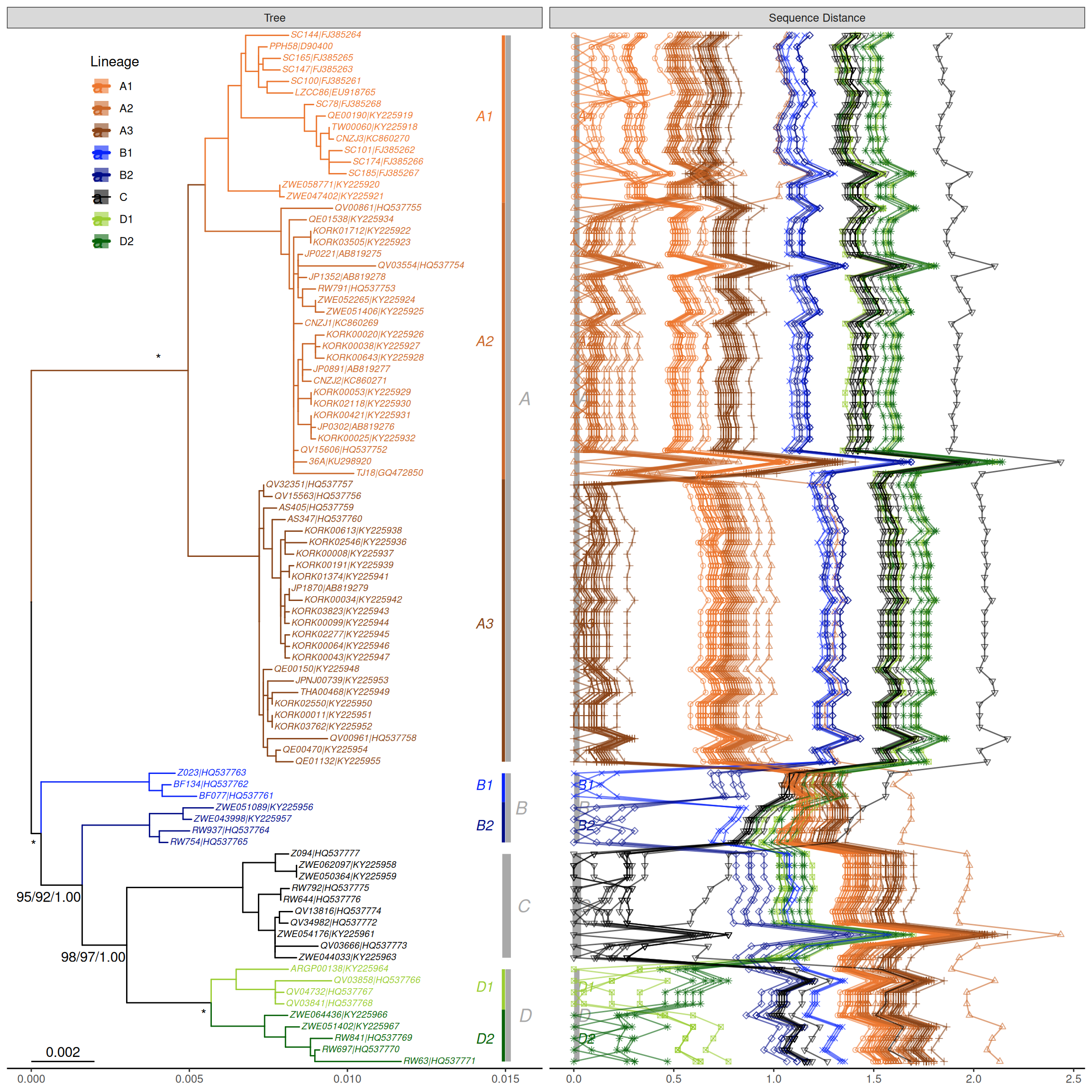
14.2 Displaying Different Symbolic Points for Bootstrap Values.
We can cut the bootstrap values into several intervals, e.g., to indicate whether the clade is of high, moderate, or low support. Then we can use these intervals as categorical variables to set different colors or shapes of symbolic points to indicate the bootstrap values belong to which category (Figure Figure 14.2).
## phytools also have a read.newick function
read.newick <- treeio::read.newicklibrary(treeio)
library(ggplot2)
library(ggtree)
library(TDbook)
tree <- read.newick(text=text_RMI_tree, node.label = "support")
root <- rootnode(tree)
ggtree(tree, color="black", size=1.5, linetype=1, right=TRUE) +
geom_tiplab(size=4.5, hjust = -0.060, fontface="bold") + xlim(0, 0.09) +
geom_point2(aes(subset=!isTip & node != root,
fill=cut(support, c(0, 700, 900, 1000))),
shape=21, size=4) +
theme_tree(legend.position=c(0.2, 0.2)) +
scale_fill_manual(values=c("white", "grey", "black"), guide='legend',
name='Bootstrap Percentage(BP)',
breaks=c('(900,1e+03]', '(700,900]', '(0,700]'),
labels=expression(BP>=90,70 <= BP * " < 90", BP < 70))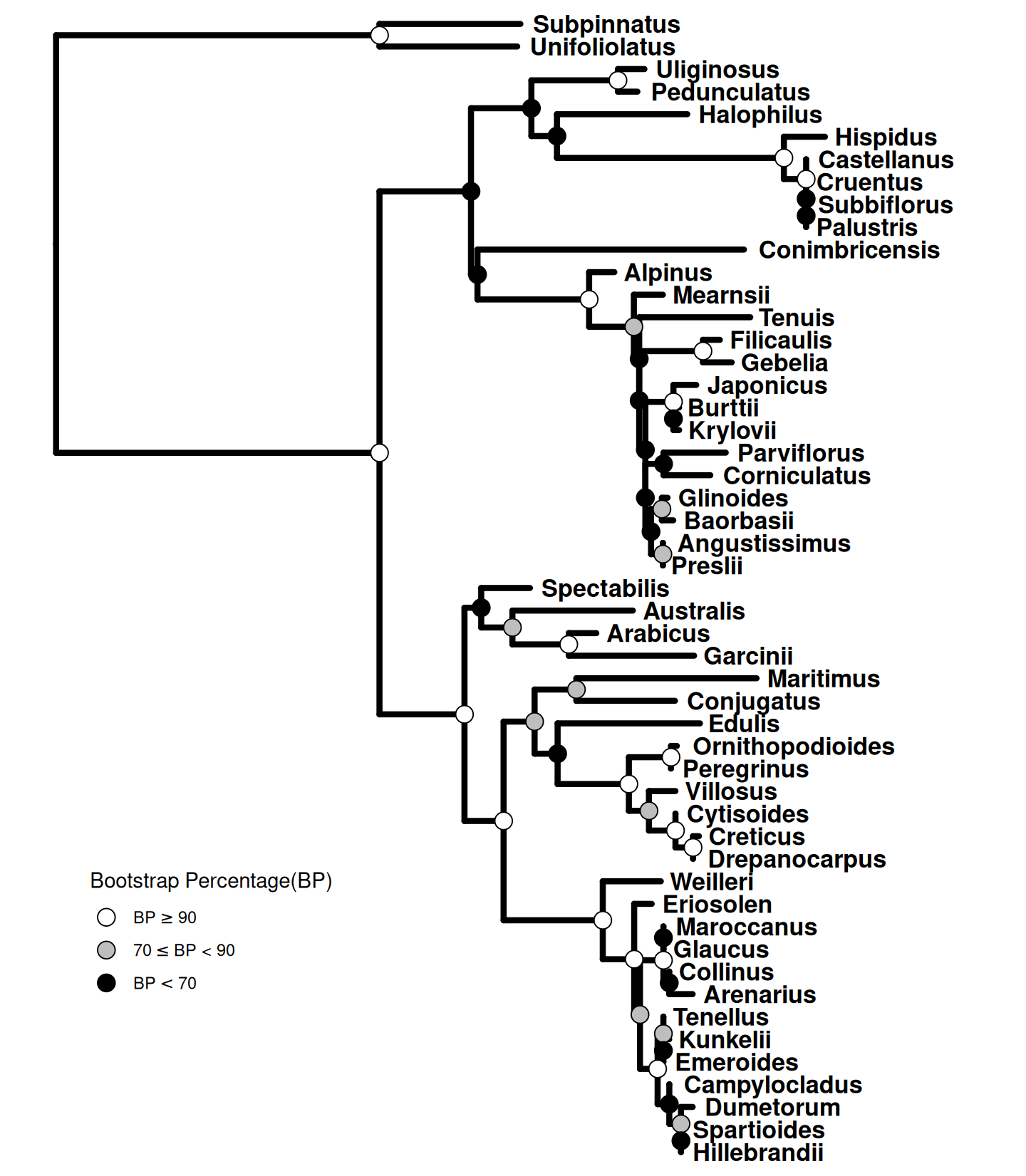
14.3 Highlighting Different Groups
This example reproduces Figure 1 of (Larsen et al., 2019). It used groupOTU() to add grouping information of chicken CTLDcps. The branch line type and color are defined based on this grouping information. Two groups of CTLDcps are highlighted in different background colors using geom_hilight (red for Group II and green for Group V). The avian-specific expansion of Group V with the subgroups of A and B- are labeled using geom_cladelab (Figure Figure 14.3).
library(TDbook)
mytree <- tree_treenwk_30.4.19
# Define nodes for coloring later on
tiplab <- mytree$tip.label
cls <- tiplab[grep("^ch", tiplab)]
labeltree <- groupOTU(mytree, cls)
p <- ggtree(labeltree, aes(color=group, linetype=group), layout="circular") +
scale_color_manual(values = c("#efad29", "#63bbd4")) +
geom_nodepoint(color="black", size=0.1) +
geom_tiplab(size=2, color="black")
p2 <- flip(p, 136, 110) %>%
flip(141, 145) %>%
rotate(141) %>%
rotate(142) %>%
rotate(160) %>%
rotate(164) %>%
rotate(131)
### Group V and II coloring
dat <- data.frame(
node = c(110, 88, 156,136),
fill = c("#229f8a", "#229f8a", "#229f8a", "#f9311f")
)
p3 <- p2 +
geom_hilight(
data = dat,
mapping = aes(
node = node,
fill = I(fill)
),
alpha = 0.2,
extendto = 1.4
)
### Putting on a label on the avian specific expansion
p4 <- p3 +
geom_cladelab(
node = 113,
label = "Avian-specific expansion",
align = TRUE,
angle = -35,
offset.text = 0.05,
hjust = "center",
fontsize = 2,
offset = .2,
barsize = .2
)
### Adding the bootstrap values with subset used to remove all bootstraps < 50
p5 <- p4 +
geom_nodelab(
mapping = aes(
x = branch,
label = label,
subset = !is.na(as.numeric(label)) & as.numeric(label) > 50
),
size = 2,
color = "black",
nudge_y = 0.6
)
### Putting labels on the subgroups
p6 <- p5 +
geom_cladelab(
data = data.frame(
node = c(114, 121),
name = c("Subgroup A", "Subgroup B")
),
mapping = aes(
node = node,
label = name
),
align = TRUE,
offset = .05,
offset.text = .03,
hjust = "center",
barsize = .2,
fontsize = 2,
angle = "auto",
horizontal = FALSE
) +
theme(
legend.position = "none",
plot.margin = grid::unit(c(-15, -15, -15, -15), "mm")
)
print(p6)Warning in FUN(X[[i]], ...): NAs introduced by coercion
Warning in FUN(X[[i]], ...): NAs introduced by coercion
Warning in FUN(X[[i]], ...): NAs introduced by coercion
Warning in FUN(X[[i]], ...): NAs introduced by coercion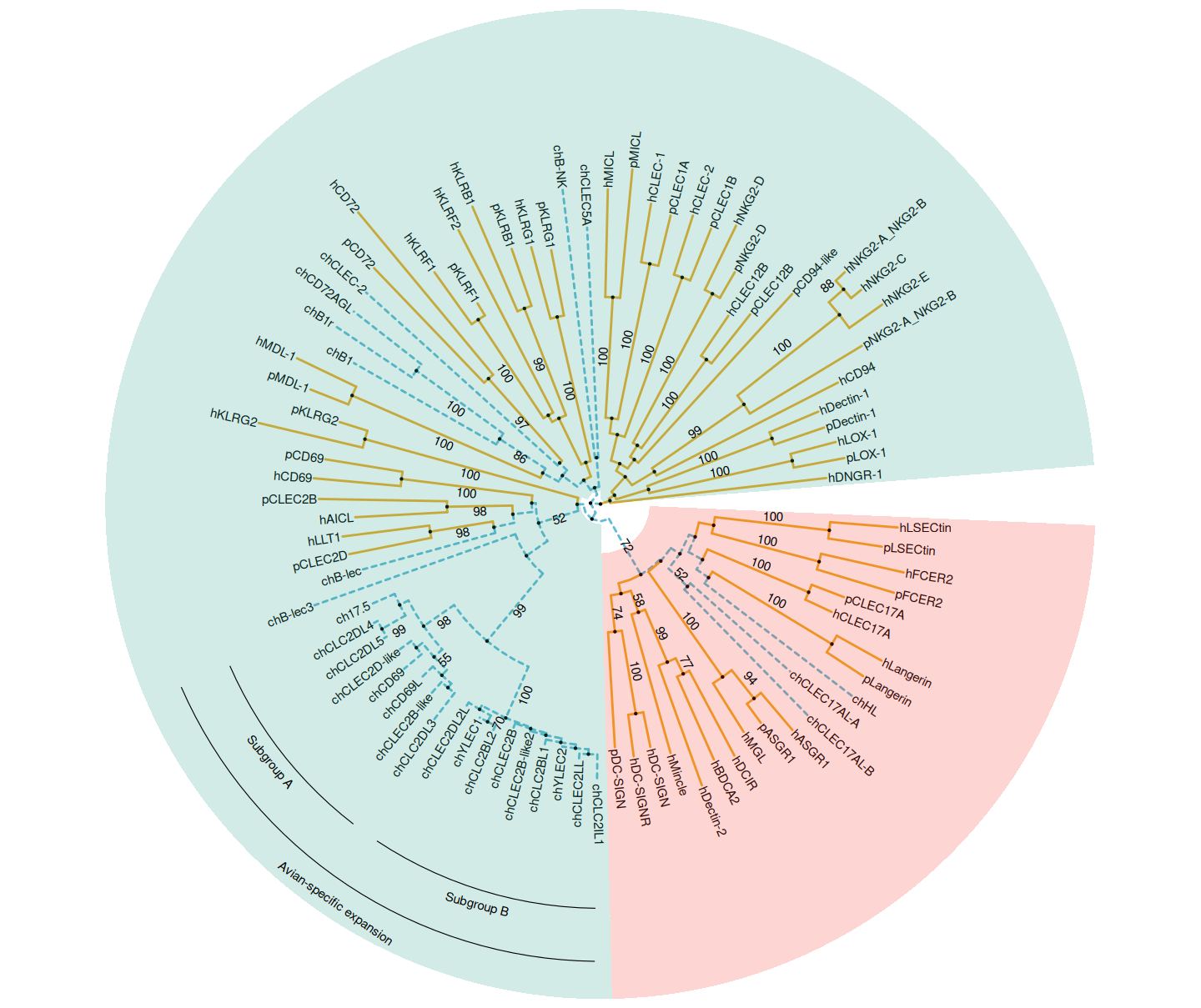
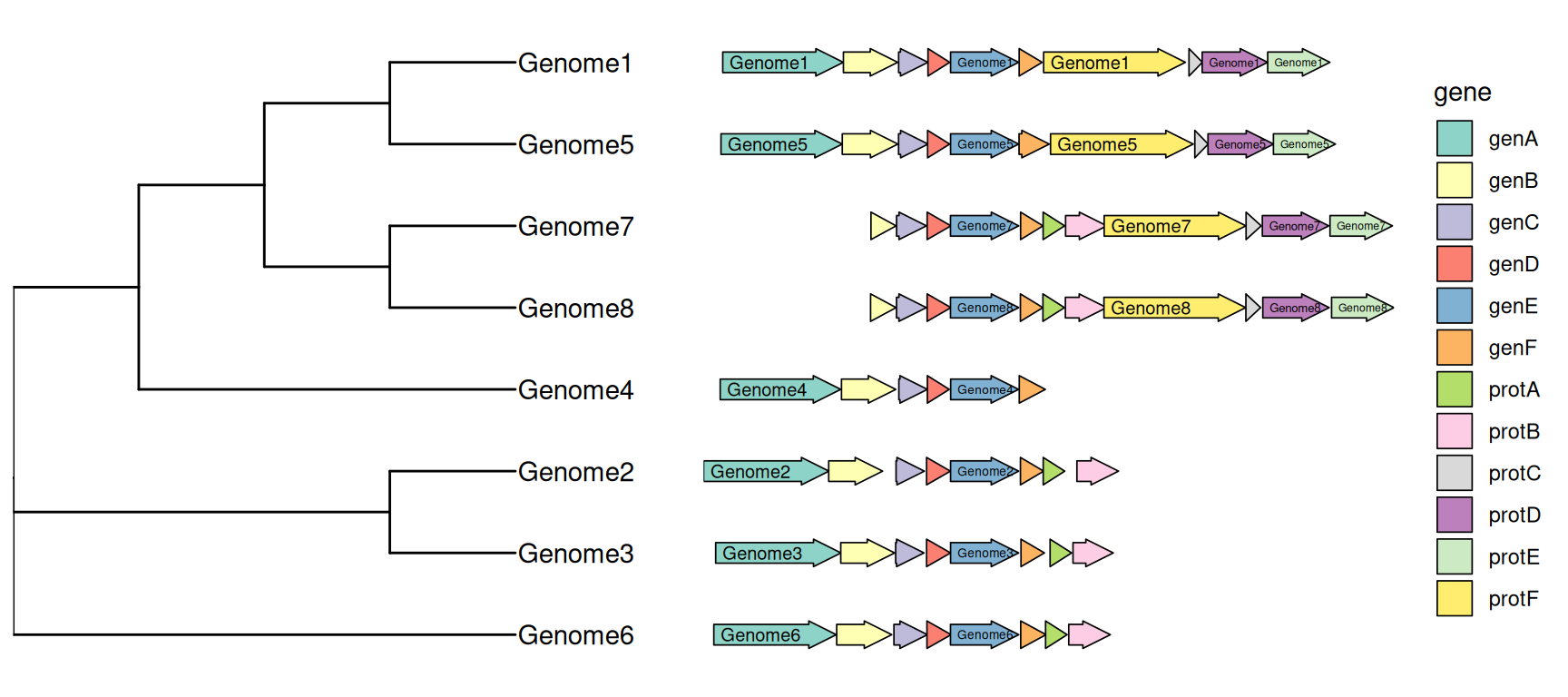
14.4 Phylogenetic Tree with Genome Locus Structure
The geom_motif() is defined in ggtree and it is a wrapper layer of the gggenes::geom_gene_arrow(). The geom_motif() can automatically adjust genomic alignment by selective gene (via the on parameter) and can label genes via the label parameter. In the following example, we use example_genes dataset provided by gggenes. As the dataset only provides genomic coordination of a set of genes, a phylogeny for the genomes needs to be constructed first. We calculate Jaccard similarity based on the ratio of overlapping genes among genomes and correspondingly determine genome distance. The BioNJ algorithm was applied to construct the tree. Then we can use geom_facet() to visualize the tree with the genomic structures (Figure Figure 14.4).
library(dplyr)
Attaching package: 'dplyr'The following objects are masked from 'package:Biostrings':
collapse, intersect, setdiff, setequal, unionThe following object is masked from 'package:GenomeInfoDb':
intersectThe following object is masked from 'package:XVector':
sliceThe following objects are masked from 'package:IRanges':
collapse, desc, intersect, setdiff, slice, unionThe following objects are masked from 'package:S4Vectors':
first, intersect, rename, setdiff, setequal, unionThe following objects are masked from 'package:BiocGenerics':
combine, intersect, setdiff, unionThe following objects are masked from 'package:stats':
filter, lagThe following objects are masked from 'package:base':
intersect, setdiff, setequal, unionlibrary(ggplot2)
library(gggenes)
library(ggtree)
get_genes <- function(data, genome) {
filter(data, molecule == genome) %>% pull(gene)
}
g <- unique(example_genes[,1])
n <- length(g)
d <- matrix(nrow = n, ncol = n)
rownames(d) <- colnames(d) <- g
genes <- lapply(g, get_genes, data = example_genes)
for (i in 1:n) {
for (j in 1:i) {
jaccard_sim <- length(intersect(genes[[i]], genes[[j]])) /
length(union(genes[[i]], genes[[j]]))
d[j, i] <- d[i, j] <- 1 - jaccard_sim
}
}
tree <- ape::bionj(d)
p <- ggtree(tree, branch.length='none') +
geom_tiplab() + xlim_tree(5.5) +
geom_facet(mapping = aes(xmin = start, xmax = end, fill = gene),
data = example_genes, geom = geom_motif, panel = 'Alignment',
on = 'genE', label = 'gene', align = 'left') +
scale_fill_brewer(palette = "Set3") +
scale_x_continuous(expand=c(0,0)) +
theme(strip.text=element_blank(),
panel.spacing=unit(0, 'cm'))
facet_widths(p, widths=c(1,2))Warning in x[i] <- value: number of items to replace is not a multiple of
replacement length