ggmsa
ggmsa.RdPlot multiple sequence alignment using ggplot2 with multiple color schemes supported.
ggmsa( msa, start = NULL, end = NULL, font = "helvetical", color = "Chemistry_AA", custom_color = NULL, char_width = 0.9, none_bg = FALSE, by_conservation = FALSE, posHighligthed = NULL, seq_name = NULL, border = NULL, consensus_views = FALSE, use_dot = FALSE, disagreement = TRUE, ignore_gaps = FALSE, ref = NULL, show.legend = FALSE )
Arguments
| msa | Multiple aligned sequence files or objects representing either nucleotide sequences or AA sequences. |
|---|---|
| start | a numeric vector. Start position to plot. |
| end | a numeric vector. End position to plot. |
| font | font families, possible values are 'helvetical', 'mono', and 'DroidSansMono', 'TimesNewRoman'. Defaults is 'helvetical'. If font = NULL, only plot the background tile. |
| color | a Color scheme. One of 'Clustal', 'Chemistry_AA', 'Shapely_AA', 'Zappo_AA', 'Taylor_AA', 'LETTER', 'CN6', 'Chemistry_NT', 'Shapely_NT', 'Zappo_NT', 'Taylor_NT'. Defaults is 'Chemistry_AA'. |
| custom_color | A data frame with two cloumn called "names" and "color".Customize the color scheme. |
| char_width | a numeric vector. Specifying the character width in the range of 0 to 1. Defaults is 0.9. |
| none_bg | a logical value indicating whether background should be disaplayed. Defaults is FALSE. |
| by_conservation | a logical value. The most conserved regions have the brightest colors. |
| posHighligthed | A numeric vector of the position that need to be highlighted. |
| seq_name | a logical value indicating whether seqence names should be displayed. Defaults is 'NULL' which indicates that the sequence name is displayed when 'font = null', but 'font = char' will not be displayed. If 'seq_name = TRUE' the sequence name will be displayed in any case. If 'seq_name = FALSE' the sequence name will not be displayed under any circumstances. |
| border | a character string. The border color. |
| consensus_views | a logical value that opeaning consensus views. |
| use_dot | a logical value. Displays characters as dots instead of fading their color in the consensus view. |
| disagreement | a logical value. Displays characters that disagreememt to consensus(excludes ambiguous disagreements). |
| ignore_gaps | a logical value. When selected TRUE, gaps in column are treated as if that row didn't exist. |
| ref | a character string. Specifying the reference sequence which should be one of input sequences when 'consensus_views' is TRUE. |
| show.legend | logical. Should this layer be included in the legends? |
Value
ggplot object
Author
Guangchuang Yu
Examples
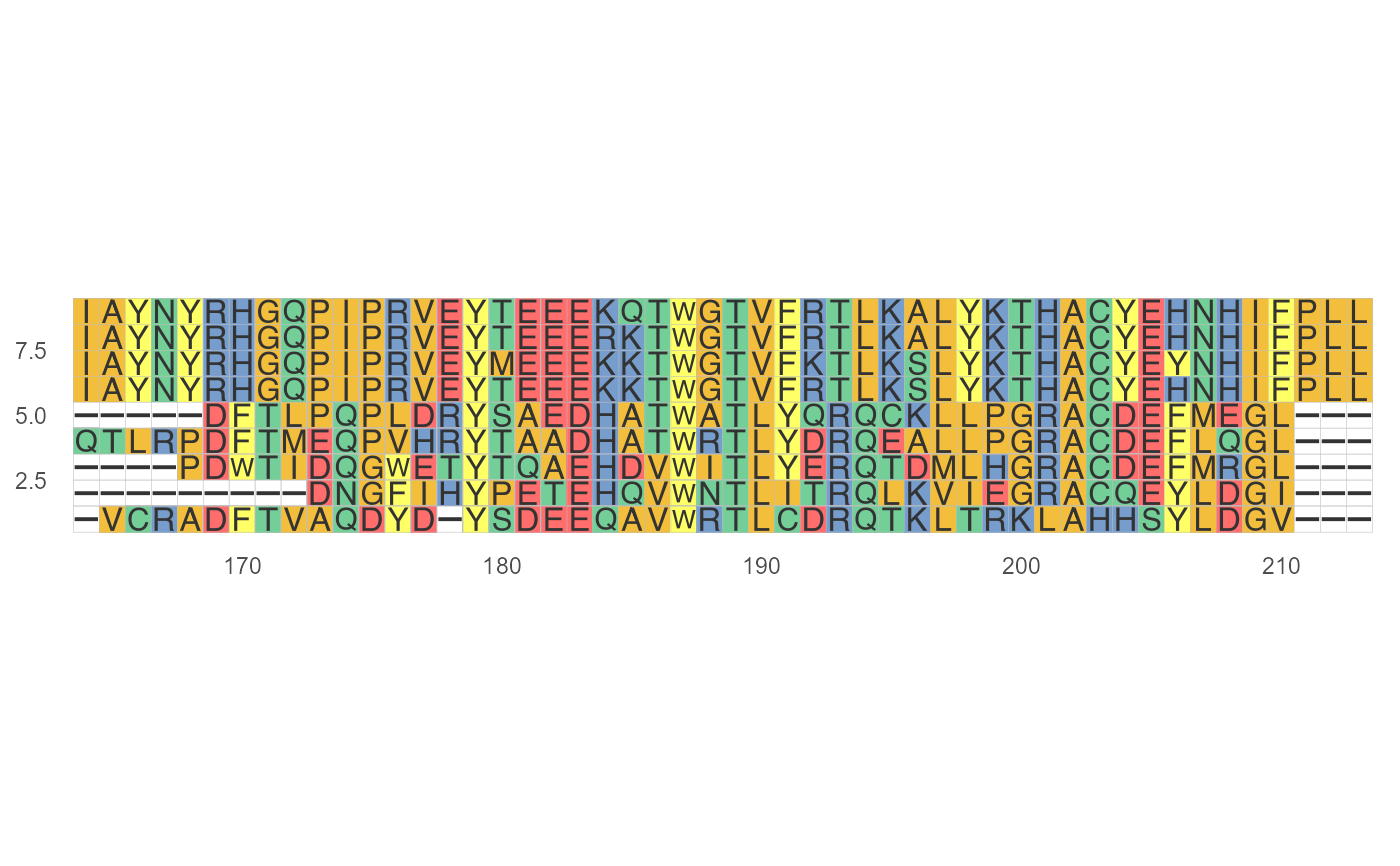
#plot multiple sequences by loading fasta format fasta <- system.file("extdata", "sample.fasta", package = "ggmsa") ggmsa(fasta, 164, 213, color="Chemistry_AA")#XMultipleAlignment objects can be used as input in the 'ggmsa' #AAMultipleAlignment <- Biostrings::readAAMultipleAlignment(fasta) #ggmsa(AAMultipleAlignment, 164, 213, color="Chemistry_AA") #XStringSet objects can be used as input in the 'ggmsa' #AAStringSet <- Biostrings::readAAStringSet(fasta) #ggmsa(AAStringSet, 164, 213, color="Chemistry_AA") #Xbin objects from 'seqmagick' can be used as input in the 'ggmsa' #AAbin <- seqmagick::fa_read(fasta) #ggmsa(AAbin, 164, 213, color="Chemistry_AA")