Dear clusterProfiler maintainers,
First, thank you for developing and maintaining this incredibly useful package! I’ve been using clusterProfiler extensively for enrichment analysis and it’s been invaluable for my research.
I’m encountering an issue with KEGG enrichment analysis where some pathways show NA values for both category and subcategory fields, even though the pathways themselves are valid and statistically significant.
Description
When performing KEGG enrichment analysis using enrichKEGG followed by setReadable, some pathways show NA values for both category and subcategory fields, even though the pathways themselves are valid and significant.
Problem
The pathways mmu04082 (Neuroactive ligand-receptor interaction) and mmu04517 (IgSF CAM signaling) return NA for category and subcategory columns in the results, while other pathways have proper classification information.
Questions
Is this a known issue with specific KEGG pathways?
Is there a way to automatically retrieve the missing category information?
Should the category information be populated from a different source or database?
I’ve verified that these pathways exist in the KEGG database and are valid, but the classification metadata seems to be missing in the enrichment results.
Any guidance on how to resolve this would be greatly appreciated. Thank you for your time and continued support of this excellent package!
Best regards,
Sophia
formula <- compareCluster(ENTREZID~cluster, data=GeneClusterDF,
fun='enrichKEGG',
organism = "mmu", # 小鼠的KEGG organism code
pvalueCutoff=0.05,
pAdjustMethod = "BH")
formula <- setReadable(formula,
OrgDb = "org.Mm.eg.db",
keyType = "ENTREZID")
head(as.data.frame(formula))
> head(as.data.frame(formula))
Cluster cluster category subcategory ID Description GeneRatio BgRatio RichFactor FoldEnrichment
1 0 0 <NA> <NA> mmu04082 Neuroactive ligand signaling 14/113 196/10632 0.07142857 6.720607
2 0 0 <NA> <NA> mmu04517 IgSF CAM signaling 16/113 300/10632 0.05333333 5.018053
3 0 0 Organismal Systems Nervous system mmu04724 Glutamatergic synapse 10/113 117/10632 0.08547009 8.041752
4 0 0 Environmental Information Processing Signal transduction mmu04020 Calcium signaling pathway 13/113 255/10632 0.05098039 4.796668
5 0 0 Organismal Systems Nervous system mmu04727 GABAergic synapse 8/113 91/10632 0.08791209 8.271516
6 0 0 Environmental Information Processing Signal transduction mmu04024 cAMP signaling pathway 11/113 224/10632 0.04910714 4.620417
zScore pvalue p.adjust qvalue geneID Count
1 8.378035 1.822353e-08 4.355424e-06 3.356966e-06 Adcy2/Gabbr2/Gabra2/Gabrb1/Gnao1/Gria2/Grin2c/Grm3/Hrh1/Plcb1/Slc1a2/Slc1a3/Slc6a1/Slc6a11 14
2 7.316833 1.016818e-07 1.215097e-05 9.365424e-06 Actr3b/Ank2/Cables1/Cadm1/Cadm2/Cntn1/Kcnq3/Kirrel3/Lrrc4c/Mapk10/Ncam1/Nrcam/Nrp1/Srgap1/Tjp1/Vav3 16
3 7.937944 4.238134e-07 3.376380e-05 2.602363e-05 Adcy2/Glul/Gnao1/Gria2/Grin2c/Grm3/Itpr2/Plcb1/Slc1a2/Slc1a3 10
4 6.360275 2.911406e-06 1.739565e-04 1.340779e-04 Adcy2/Atp2b2/Camk1d/Camk2g/Fgf1/Fgfr3/Grin2c/Hrh1/Itpr2/Phka1/Phkg1/Plcb1/Vegfa 13
5 7.220110 5.181671e-06 2.476839e-04 1.909037e-04 Adcy2/Gabbr2/Gabra2/Gabrb1/Glul/Gnao1/Slc6a1/Slc6a11 8
6 5.675943 2.505014e-05 9.978306e-04 7.690833e-04 Adcy2/Atp1a2/Atp2b2/Camk2g/Gabbr2/Gli3/Gria2/Grin2c/Mapk10/Ptch1/Vav3 11
> sessionInfo()
R version 4.5.2 (2025-10-31 ucrt)
Platform: x86_64-w64-mingw32/x64
Running under: Windows 11 x64 (build 22000)
Matrix products: default
LAPACK version 3.12.1
locale:
[1] LC_COLLATE=Chinese (Simplified)_China.utf8 LC_CTYPE=Chinese (Simplified)_China.utf8 LC_MONETARY=Chinese (Simplified)_China.utf8
[4] LC_NUMERIC=C LC_TIME=Chinese (Simplified)_China.utf8
time zone: Asia/Shanghai
tzcode source: internal
attached base packages:
[1] stats4 stats graphics grDevices utils datasets methods base
other attached packages:
[1] dplyr_1.1.4 ReactomePA_1.54.0 GseaVis_0.0.5 pathview_1.50.0 DOSE_4.4.0 enrichplot_1.30.0
[7] ggplot2_4.0.1 msigdbr_25.1.1 org.Mm.eg.db_3.22.0 AnnotationDbi_1.72.0 IRanges_2.44.0 S4Vectors_0.48.0
[13] Biobase_2.70.0 BiocGenerics_0.56.0 generics_0.1.4 clusterProfiler_4.18.0
loaded via a namespace (and not attached):
[1] pheatmap_1.0.13 pak_0.9.0 DBI_1.2.3 httr_1.4.7 BiocParallel_1.44.0 yulab.utils_0.2.1
[7] ggplotify_0.1.3 babelgene_22.9 pillar_1.11.1 Rgraphviz_2.54.0 R6_2.6.1 mime_0.13
[13] reticulate_1.44.1 viridis_0.6.5 ROCR_1.0-11 graphite_1.56.0 S7_0.2.1 parallelly_1.45.1
[19] GlobalOptions_0.1.2 polyclip_1.10-7 htmltools_0.5.8.1 remotes_2.5.0 ggrepel_0.9.6 fgsea_1.36.0
[25] forcats_1.0.1 spatstat.utils_3.2-0 fitdistrplus_1.2-4 tidyselect_1.2.1 utf8_1.2.6 RSQLite_2.4.4
[31] cowplot_1.2.0 scattermore_1.2 sessioninfo_1.2.3 spatstat.data_3.1-9 gridExtra_2.3 fs_1.6.6
[37] sctransform_0.4.2 RColorBrewer_1.1-3 future.apply_1.20.0 graph_1.88.0 R.oo_1.27.1 RcppHNSW_0.6.0
[43] reactome.db_1.94.0 Rtsne_0.17 lazyeval_0.2.2 scales_1.4.0 treeio_1.34.0 R.utils_2.13.0
[49] KEGGgraph_1.70.0 bitops_1.0-9 R.methodsS3_1.8.2 KEGGREST_1.50.0 promises_1.5.0 shape_1.4.6.1
[55] zoo_1.8-14 RSpectra_0.16-2 assertthat_0.2.1 tools_4.5.2 ape_5.8-1 shiny_1.11.1
[61] rlang_1.1.6 ggridges_0.5.7 evaluate_1.0.5 otel_0.2.0 reshape2_1.4.5 devtools_2.4.6
[67] colorspace_2.1-2 ellipsis_0.3.2 data.table_1.17.8 withr_3.0.2 tibble_3.3.0 RCurl_1.98-1.17
[73] xtable_1.8-4 plyr_1.8.9 aplot_0.2.9 systemfonts_1.3.1 httpuv_1.6.16 MASS_7.3-65
[79] stringr_1.6.0 openxlsx_4.2.8.1 GO.db_3.22.0 vctrs_0.6.5 lifecycle_1.0.4 codetools_0.2-20
[85] fastDummies_1.7.5 nlme_3.1-168 Seqinfo_1.0.0 future_1.68.0 pkgload_1.4.1 Rcpp_1.1.0
[91] rstudioapi_0.17.1 patchwork_1.3.2 stringi_1.8.7 pbapply_1.7-4 cachem_1.1.0 BiocManager_1.30.27
[97] tidytree_0.4.6 listenv_0.10.0 XVector_0.50.0 plotly_4.11.0 ggtree_4.0.1 pkgbuild_1.4.8
[103] ggfun_0.2.0 ggtangle_0.0.8 htmlwidgets_1.6.4 memoise_2.0.1 crayon_1.5.3 gridGraphics_0.5-1
[109] rappdirs_0.3.3 GOSemSim_2.36.0 png_0.1-8 progressr_0.18.0 fastmap_1.2.0 tidygraph_1.3.1
[115] tidyr_1.3.1 pkgconfig_2.0.3 cli_3.6.5 ggforce_0.5.0 ggiraph_0.9.2 lmtest_0.9-40
[121] usethis_3.2.1 RcppAnnoy_0.0.22 gdtools_0.4.4 viridisLite_0.4.2 splines_4.5.2 blob_1.2.4
[127] XML_3.99-0.20 globals_0.18.0 knitr_1.50 ica_1.0-3 spam_2.11-1 dichromat_2.0-0.1
[133] compiler_4.5.2 grid_4.5.2 bit_4.6.0 ggpp_0.5.9 glue_1.8.0 sp_2.2-0
[139] digest_0.6.38 irlba_2.3.5.1 graphlayouts_1.2.2 fontLiberation_0.1.0 fontBitstreamVera_0.1.1 dotCall64_1.2
[145] tweenr_2.0.3 lattice_0.22-7 ggraph_2.2.2 gson_0.1.0 igraph_2.2.1 ggnewscale_0.5.2
[151] qvalue_2.42.0 later_1.4.4 parallel_4.5.2 fontquiver_0.2.1 miniUI_0.1.2 gtable_0.3.6
[157] xfun_0.54 Biostrings_2.78.0 curl_7.0.0 org.Hs.eg.db_3.22.0 KernSmooth_2.23-26 survival_3.8-3
[163] jsonlite_2.0.0 magrittr_2.0.4 purrr_1.2.0 matrixStats_1.5.0 Matrix_1.7-4 SeuratObject_5.2.0
[169] fastmatch_1.1-6 RANN_2.6.2 circlize_0.4.16 polynom_1.4-1 bit64_4.6.0-1 cluster_2.1.8.1
[175] farver_2.1.2 zip_2.3.3 来自用户的灵魂拷问,连着三个问句,这必须是有经历过探索的。我看了一下,这个KEGG category的映射关系是预存在包里的,然后在Bioconductor 3.22新发行版的时候,我忘记去更一下数据了。
那么KEGG本来就是在线抓取数据,为什么这个要预存数据呢?也在线爬它不香吗?这个我肯定是有原因的。
-
因为这个category,不管你是什么物种,都是这样的分类,所以它是通用的。所有人一样用,这就适合存一份。
-
另一方面,爬这个category的信息,我用的
rvest去写的爬虫,我不想放到clusterProfiler的代码里,因为放进去,就会增加依赖包。
我给用户的回复:
The KEGG pathway information is cached within the package. I apologize for not updating it before the latest release. Thank you for bringing this to my attention; the issue has now been resolved.
> head(x, 2)
category subcategory ID Description GeneRatio BgRatio RichFactor FoldEnrichment zScore pvalue p.adjust qvalue
mmu04082 Environmental Information Processing Signaling molecules and interaction mmu04082 Neuroactive ligand signaling 30/30 196/10632 0.1530612 54.24490 40.02145 9.349530e-54 1.084545e-51 4.527141e-52
mmu05032 Human Diseases Substance dependence mmu05032 Morphine addiction 13/30 93/10632 0.1397849 49.53978 25.00883 7.801411e-20 4.524818e-18 1.888763e-18
geneID Count
mmu04082 216227/11423/14678/14701/11549/14654/18750/57385/11539/15559/14683/11515/11541/104111/18442/242425/14806/13491/21334/63993/13488/213788/57014/108015/14396/21337/53623/110886/11513/210044 30
mmu05032 昨晚给我的留言,早上就许愿成功。

这个锅我是认的。
正如前文所的,预存数据是有原因的,但这也带来了要定期更新的问题。因为这才更没多久,又来反映没更了。这样子手工更,就太费叔了,一下子把自己搞手残了。
所谓那里有压迫,那里就有反抗,不能让它把我们给欺负了。
我的解决方案就是用github action。写个workflow来干这个事情。
- 触发条件 :每周日 UTC 时间 00:00 ( cron: ‘0 0 * * 0’ ) 或手动触发 ( workflow_dispatch )。
- 环境配置 :自动安装 R 环境和必要的依赖(包括 rvest , tibble , tidyr 以及 clusterProfiler 自身)。
- 执行更新 :安装当前包后,完全按照我写好的 make updatedata 逻辑运行。
- 自动 PR :如果有文件变动( .rda 文件更新),自动创建一个名为 update-kegg-data 的分支并提交 PR。
我手动触发,试一下它work不work。显然是没有问题的,pull request已经正常出现。那么我只要merge就行了。做到数据每周定期检查,如有更新,就自动更新。
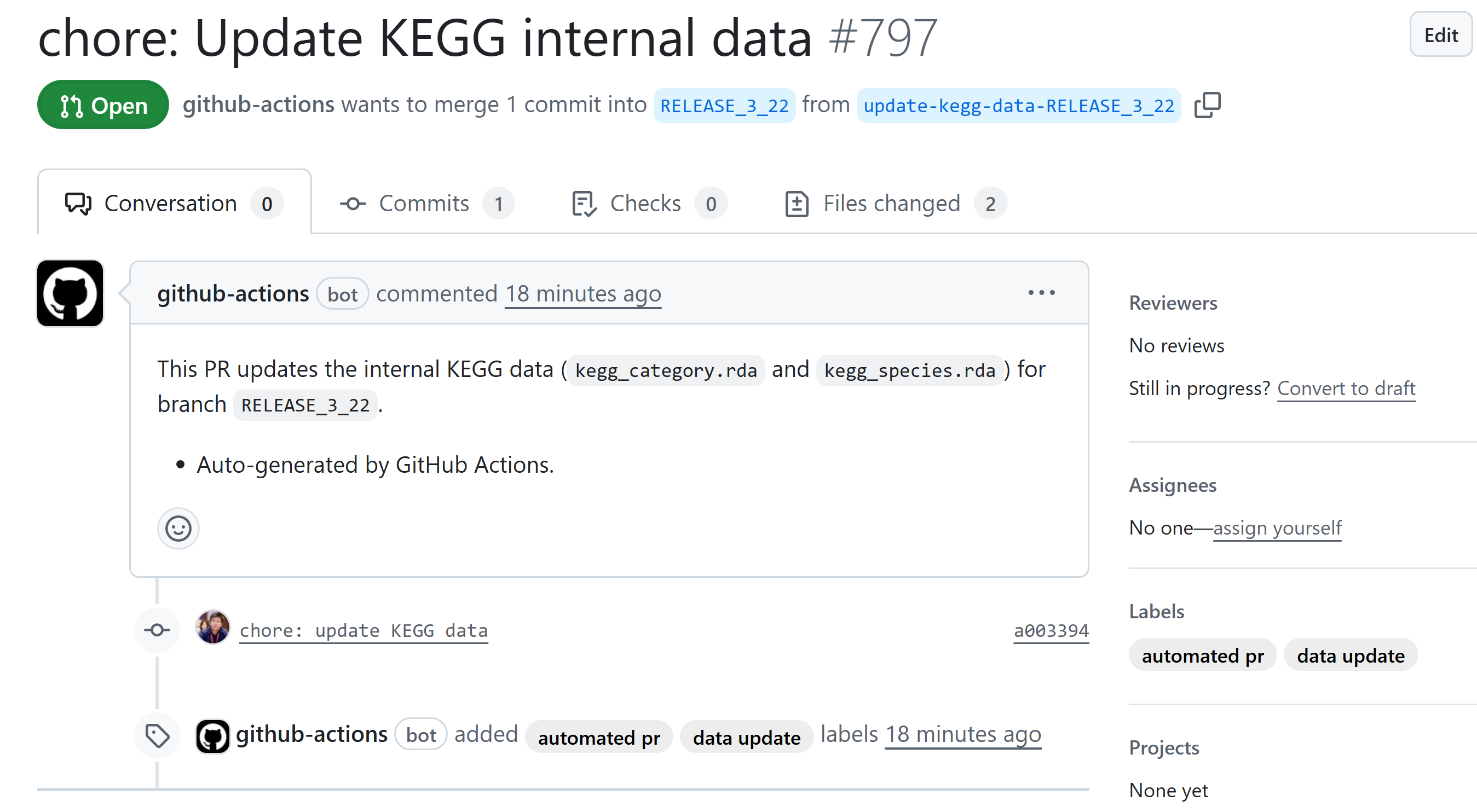 好了,活人终究不能被尿憋死。
好了,活人终究不能被尿憋死。
后续
终究我还是懒,KEGG更新还挺频繁的,我不想手工merge,再push到Bioconductor。于是数据和包分离,数据是预存没错,但不放在包里,而是放到github上,然后GitHub Actions自动更新。包里不带数据,总是从github上获取,这样最终可以做到不用管。